../Ancestry_DNA_Tests/Ancestry_DNA_Tests.htm
The first anatomically modern humans originated in Africa about 200,000 years ago, and all humans today are their direct descendants. The study points to an area along the Namibia-South Africa border, the homeland of the San people, as the starting point for a southwest-to-northeast migratory route that carried people through Africa and across the Red Sea into Eurasia.
Africans are more genetically diverse than the inhabitants of the rest of the world combined, according to a sweeping study that carried researchers into remote regions to sample the bloodlines of more than 100 distinct populations. So says Sarah Tishkoff, a University of Pennsylvania geneticist who led the international research team. The report was published in the journal Science Express.
 |
Genetic data corroborates the mitochondrial results, placing the root of the human family tree - our most recent common ancestor- in Africa within the past few hundred thousand years. Consistent with this result, all of the genetic data shows the greatest number of polymorphisms in Africa - there is simply far more variation in that continent than anywhere else. You are more likely to sample extremely divergent genetic lineages within a single African village than you are in whole of the rest of the world. The majority of the genetic polymorphisms found in our species are found uniquely in Africans - Europeans, Asians and Native Americans carry only a small sample of the extraordinary diversity that can be found in any African village.
Our Human GenesEach gene resides at a specific locus (location on a chromosome) in two copies, one copy of the gene inherited from each parent. As a simplistic example: When two Chinese mate, the child will look Chinese because all the genes are healthy and all the genes are the same. But if a Chinese and a White European mate, the children will look like some combination of the two, because the "Appearance" genes are not all the same. Gene copies, however, are not always healthy. When the copies of a gene differ from each other, as through deleterious mutation or failure: Then in this heterozygous condition, we call the two parts “Alleles” and the undamaged or un-mutated allele is dominant, and the organism’s appearance and function is normal. The damaged "other" allele has no noticeable effect on the organism’s appearance, and is called the “Recessive” allele. When BOTH alleles of a gene become recessive, then the gene cannot complete its assignment. As an example: many Black people have alleles of their “P” gene which are heterozygous and they look normal in every way: (The “P” gene controls the production of Melanin in the skin for protection from the Sun). But if TWO of these people with heterozygous alleles in their “P” gene MATE, then one or more, of their children will be an Albino. If two Albinos mate, there is only damaged or recessive “P” genes to inherit; therefore ALL of their children will be White. The trait for curly hair (which is the normal for humans) follows the same rules, two damaged or recessive allele’s of the "TCHH" gene means straight hair. Same for the genes which control eye color and hair color: (Blonde and Red hair is recessive, as is Blue, Green, and Gray eyes). Note: The trait for Curly/Kinky hair (which is the "Normal" for humans): is produced by two "Undamaged" TCHH genes. That means that "Curly Hair" is "ANCESTRAL" to Modern Humans. You might keep that in mind the next time you see the White mans depictions of ancient humans shown with "Straight" hair. |
Albino people try to embed three thoughts into our minds: 1) That they are unique, a completely separate branch of the Human Tree, a different "Race" if you will: and certainly NOT Albinos: 2) They ARE Native to Europe: 3) They were the "Original" people of Mans Ancient Civilizations. All of which are of course total and utter nonsense. But in trying to make those absurdities seem plausible, they create all sorts of Pseudo-Scientific Lies. The news-story below seeks to embed in us, belief in the lie that there is such a thing as "WHITE PEOPLES GENES".
 |
By Damien Gayle - Published: 16:13 EST, 16 May 2013
DNA analysis has debunked the longstanding theory that the Minoans, who some 5,000 years ago established Europe's first advanced Bronze Age culture, were from Africa. The Minoan civilisation arose on the Mediterranean island of Crete in approximately the 27th century BC and flourished for 12 centuries until the 15th century BC.But the culture was lost until British archaeologist Sir Arthur Evans unearthed its remains on Crete in 1900, where he found vestiges of a civilisation he believed was formed by refugees from northern Egypt.
Modern archaeologists have cast doubt on that version of events, and now DNA tests of Minoan remains suggests they were descended from ancient farmers who settled the islands thousands of years earlier. These people, it is believed, are from the same stock that came from the East to populate the rest of Europe. Evans set to work on Crete in 1900 with a team of archaeologists soon after the island was liberated from the yoke of the Ottoman empire, almost immediately unearthing a great palace.
He named the civilisation he discovered after the legendary Greek king Minos and, based on likenesses between Minoan artifacts and those from Egypt and Libya, proposed that its founders migrated into the area from North Africa. Since then, other archaeologists have suggested that the Minoans may have come from other regions, possibly Turkey, the Balkans, or the Middle East. But now a joint U.S. and Greek team has made a mitochondrial DNA analysis of Minoan skeletal remains to determine the likely ancestors of the ancient people. Mitochondria, the energy powerhouses of cells, contain their own DNA, or genetic code, and because mitochondrial DNA is passed down from mothers to their children via the human egg, it contains information about maternal ancestry.
Findings suggest that the Minoan civilisation arose from the population already living in Crete, and that these people were probably descendants of the first humans to reach there about 9,000 years ago. Further, they found, the remains have the greatest genetic similarity with modern European populations. Senior researcher Dr George Stamatoyannopoulos, professor of medicine and genome sciences at the University of Washington, said the analysis showed these people probably came to the area from the East, not the South.
'About 9,000 years ago there was an extensive migration of Neolithic humans from the regions of Anatolia that today comprise parts of Turkey and the Middle East,' he said.'At the same time, the first Neolithic inhabitants reached Crete. 'Our mitochondrial DNA analysis shows that the Minoans' strongest genetic relationships are with these Neolithic humans, as well as with ancient and modern Europeans. 'These results suggest the Minoan civilization arose 5,000 years ago in Crete from an ancestral Neolithic population that had arrived in the region about 4,000 years earlier. 'Our data suggest that the Neolithic population that gave rise to the Minoans also migrated into Europe and gave rise to modern European peoples.'
Dr Stamatoyannopoulos and his team analysed samples from 37 skeletons found in a cave in Crete’s Lassithi plateau and compared them with mitochondrial DNA sequences from 135 modern and ancient human populations. The Minoan samples revealed 21 distinct mitochondrial DNA variations, of which six were unique to the Minoans and 15 were shared with modern and ancient populations. None of the Minoans carried mitochondrial DNA variations characteristic of African populations. Further analysis showed that the Minoans were only distantly related to Egyptian, Libyan, and other North African populations. Indeed, the Minoan shared the greatest percentage of their mitochondrial DNA variation with European populations, especially those in Northern and Western Europe.
When plotted geographically, shared Minoan mitochondrial DNA variation was lowest in North Africa and increased progressively across the Middle East, Caucasus, Mediterranean islands, Southern Europe, and mainland Europe. The highest percentage of shared Minoan mitochondrial DNA variation was found with Neolithic populations from Southern Europe. The analysis also showed a high degree of sharing with the current population of the Lassithi plateau and Greece. In fact, the maternal genetic information passed down through many generations of mitochondria is still present in modern-day residents of the area where the Minoan skeletons were found. Dr Stamatoyannopoulos said he believes that the findings highlight the importance of DNA analysis as a tool for understanding human history.
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
The New York Review of Books
Knossos: Fakes, Facts, and Mystery
by Mary Beard, August 13, 2009 Issue
The masterpieces of Minoan art are not what they seem. The vivid frescoes that once decorated the walls of the prehistoric palace at Knossos in Crete are now the main attraction of the Archaeological Museum in the modern city of Heraklion, a few miles from the site of Knossos. Dating from the early or mid-second millennium BC, they are some of the most famous icons of ancient European culture, reproduced on countless postcards and posters, T-shirts and refrigerator magnets: the magnificent young “prince” with his floral crown, walking through a field of lilies; the five blue dolphins patrolling their underwater world between minnows and sea urchins; the three “ladies in blue” (a favorite Minoan color) with their curling black hair, low-cut dresses, and gesticulating hands, as if they have been caught in mid-conversation. The prehistoric world they evoke seems in some ways distant and strange—yet, at the same time, reassuringly recognizable and almost modern.
The truth is that these famous icons are largely modern. As any sharp-eyed visitor to the Heraklion museum can spot, what survives of the original paintings amounts in most cases to no more than a few square inches. The rest is more or less imaginative reconstruction, commissioned in the first half of the twentieth century by Sir Arthur Evans, the British excavator of the palace of Knossos (and the man who coined the term “Minoan” for this prehistoric Cretan civilization, after the mythical King Minos who is said to have held the throne there). As a general rule of thumb, the more famous the image now is, the less of it is actually ancient.
A quick explanation of how genetic type is passed down through generations.Europeans are Dravidian Albinos from Central Asia (that is why they and the Black Dravidians, share the same DNA); they invaded Europe twice: the first time was about 1,200 B.C. When the dust had settled hundreds of years later, they had been absorbed into the originally Black, Greek and Roman populations. Which became mixed race populations, as their artifacts clearly indicate. The second invasion was at the beginning of the current era (A.D.). The predominate Albino people were people whom we call "the Germanics". It is clear from the writings of the Roman historian Tacitus, that as the Germanics moved into new territories, the indigenous Black Europeans, were killing German Males, and taking their Females as spoils of War. Thus their offspring gained the ability to produce "Some" Melanin in their skin, and the Males gained a strengthening measure of genetic diversity. But most importantly, the German females were not taken as wives, they were simply "despoiled" and allowed to return to their tribes. Y-dna does not change, it is passed from father to son, regardless of whether the father is Black or White. Thus their "Mulatto" Male offspring would retain the Y-dna haplogroup "I or R" of their Black despoiler father. When these mulatto males bred with their tribal White females, their resultant male offspring would be Quadroons (1/4) Black, but still with the Y-dna haplogroup "I or R" of their despoiler grandfather. When these Quadroon males bred with their tribal White females, their resultant male offspring would be Octoroons (1/8) Black, but still with the Y-dna haplogroup "I or R" of their despoiler great grandfather - and so on. Blacks were the original settlers of Europe, some them, especially in Central Europe, were also Y-dna haplogroup "R".
Results of the DNA tests indicate that in the Nuclear family, the Father was Y-dna R1a, the Mother was MTdna - K
|
From Wikipedia:
According to the Genographic Project conducted by the National Geographic Society, Haplogroup R2a arose about 25,000 years ago in Central Asia and its members migrated southward as part of the second major wave of human migration into India.
According to Sengupta et al. (2006),
uncertainty neutralizes previous conclusions that the intrusion of HGs R1a1 and R2 [Now R-M124] from the northwest in Dravidian-speaking southern tribes is attributable to a single recent event. Rather, these HGs contain considerable demographic complexity, as implied by their high haplotype diversity. Specifically, they could have actually arrived in southern India from a southwestern Asian source region multiple times, with some episodes considerably earlier than others.
The following is Manoukian's (2006) summary of the findings of the Genographic Project conducted by the National Geographic Society and directed by Spencer Wells (2001):
Haplogroup R, the ancestral clade to R1 and R2, appeared on the Central Asian Steppes around 35,000 to 30,000 years ago.
R1, sister clade to R2, moved to the West (READ EUROPE) from the Central Asian Steppes around 35,000 to 30,000 years ago. R1 pockets were established, from where R1a and R1b emerged.
R2a [R-M124] made its first entry into the Indian sub-continent around 25,000 years ago. The routes taken are not clear, although the Indus and Ganges rivers are possible theories put forward. There could, of course, have been multiple immigrations of this haplogroup into the Indian sub-continent, both in the Paleolithic and the Neolithic
 |
Haplogroup R1b (Y-DNA) is the dominant paternal lineage of Western Europe, so of course the Albino people want to stake out that particular Haplogroup as the "WHITE" haplogroup. But there is just one problem, there are also lots of R1b people in Africa.
 |
Here is what the Albino people say about haplogroups "R and R1b": For "R" quote - Possible place of origin Central Asia. For "R1B" quote: Possible place of origin West Asia, Russian Plain or Central Asia. Yes, you read that right, the Albinos admit that all humans originated in Africa, and from there, spread out to populate the rest of the world, but somehow, the "White Peoples" haplogroup started in Asia! For those who know that European Albinos claim to be indigenous to Europe, yet they now claim that their genes originated in Asia: please stifle the laughter until this part is finished.
Sometimes it's almost embarrassing watching Albinos twist themselves into pretzels in order to get the actual facts to fit their made-up scenarios. So how did they try to explain how all of those Black Africans could have their "WHITE" genes"? Well those completely devoid of sense and honesty tried to bring forward the concept of a "Back Migration to Africa". But when they saw that it wouldn't work, they just let it quietly fade away.
Of course there have been Back migrations to Africa in the current era (A.D.) The Central Asian Albino tribes called: the Alans, the Vandals, and the Visigoth's, rampaged through Europe, and entered North Africa through Spain in the early part of the current era. They gave rise to the Mulatto people of North Africa calling themselves "The Amazigh". Later, around 600 A.D. a Central Asian Albino super-tribe called the Turks, entered West Asia as the Slave Soldiers (Mamluks) of the Black Arabs. Later they took over West Asia as the Ottoman Empire. Their Mulattoes are now the predominate people of all of West Asia.
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
Haplogroup M (mtDNA)
In human mitochondrial genetics, Haplogroup "M" is a human mitochondrial DNA (mtDNA) haplogroup. An enormous haplogroup spanning all the continents, the macro-haplogroup M, like its sibling "N", is a descendant of haplogroup "L3".
Haplogroup U (mtDNA)
In human mitochondrial genetics, Haplogroup "U" is a human mitochondrial DNA (mtDNA) haplogroup. "U" subgroups are widely distributed across Western Eurasia, North Africa, and South Asia.
 |
 |
 |
 |
 |
 |
Y DNA (from the Andamanese wiki).
The male Y-chromosome in humans is inherited exclusively through paternal descent. Male Onges and Jarawas almost exclusively belong to Haplogroup D-M174. The clade is most common today in Tibet and Japan, with its highest frequencies worldwide in the Pumi population of northwestern China (70.2%). Haplogroup D-M174 also occurs frequently among the Ainu (of Japan). On the Indian mainland, it has been observed at low frequencies. Andamanese males were found to carry five different binary D haplotypes, all of which had previously been observed on the Indian subcontinent, in Southeast Asia and Melanesia.
 |
 |
 |
 |
Tutankhamun’s Blood:Why everyone from the Mormons to the Muslim Brotherhood is desperate for a piece of the Pharaoh.
BACKGROUND:
__________________________________________________________________________
This article is an abject lesson in how Albinos and their Mulattoes, hold Black property, Black achievement, and the totality of Black history, at their delusional, lying whim.
THIRTY FEET BENEATH THE DESERT of southern Egypt, Yehia Gad stands in a cramped, stone tomb. Gad moves slowly, encased in a protective mask and gown, and a hat that hides his neat, grey hair. In front of him, on a wooden table, is the body that was buried here more than 3,000 years ago. The stick-thin figure is little more than a silhouette, black as coal, with empty eye sockets and skin that’s cracked like parched earth. This is Tutankhamun, Egypt’s most famous pharaoh, a man whose people believed was a god on earth.
Now American television, with its lavish budgets, has bought its way to the king. The Discovery Channel has paid millions of dollars to film a pioneering study of Tutankhamun’s genetic heritage, this time carried out by the Egyptians themselves. If successful, the project could fill state coffers, achieve a scientific coup and reclaim dented national pride. Yet the goal is so ambitious that many of the world’s top researchers insist it isn’t even possible. Gad, one of the country’s top geneticists, was chosen to lead the team. He has just one chance to collect the stories hidden deep within the king’s crumbling bones.
Autopsies were fairly limited affairs in Carter’s day, but he and his colleagues measured the mummy’s bones and examined the body for wounds. They determined that he had died young, around the age of 18. The shape of his skull also suggested that he was closely related to an anonymous pharaoh found buried nearby, in a controversial tomb called KV55. But that was as much as they could glean. Getting to the bottom of Tutankhamun’s family history would require new technology to be invented, and new science to be discovered. Gad, a post-doctoral scientist at Egypt’s National Research Center, learned how to use PCR, and was becoming alive to its possibilities. His lab had been one of the first in Egypt to offer DNA fingerprinting, which hones in on specific regions of the genome that are known to vary between individuals, and is often used in forensics. If a trace of blood or semen was found at a crime scene, scientists could use PCR to amplify the DNA, and then use fingerprinting to identify a suspect.
|
Quote from the study: "Determination of optimal rehydration, fixation and staining methods for histological and immunohistochemical analysis of mummified soft tissues"
A-M Mekota, M Vermehren.
Department of Biology I, Biodiversity Research/Anthropology1 and Department of Veterinary Anatomy II2 , Ludwig-Maximilians University Munich, Germany.
Skin sections showed particularly good tissue preservation, although cellular outlines were never distinct. Although much of the epidermis had already separated from the dermis, the remaining epidermis often was preserved well (Fig. 1). The basal epithelial cells were packed with melanin as expected for specimens of Negroid origin.
__________________________________________________________________________
Continuing:
Finally, the team got its first result from the boy king: a snatch of Tutankhamun’s Y-chromosome. Today, Gad says he can’t remember the actual moment when they realized they had their results. The version laid down in his memory is the one that the team re-enacted later for the TV cameras: a close-up of colored peaks on a computer screen followed by smiles and cheers, and team members shaking white-gloved hands.
Gad and the team had exciting news for the waiting journalists. After amplifying DNA from every mummy they tested, they had constructed a five-generation family tree. The anonymous KV55 mummy, the team said, was actually Tutankhamun’s father, the revolutionary Akhenaten, while the fetuses were most likely his daughters. But the most jaw-dropping revelation was the secret that had felled the 18th Dynasty: Tutankhamun’s parents had been siblings.
Hawass ensured that the announcement was accompanied by a media blitz, including a research paper published in the esteemed Journal of the American Medical Association and a four-hour special on the Discovery Channel called King Tut Unwrapped. He later took to the pages of National Geographic to play up the ancient soap opera. The union between Akhenaten and his sister “planted the seed of their son’s early death,” he wrote. “Tutankhamun’s health was compromised from the moment he was conceived.”
The team didn’t publish any information on the mummies’ racial or ethnic origins, saying that the data on the issue was incomplete. But that didn’t stop others from speculating. A Swiss genealogy company named IGENEA issued a press release based on a blurry screen-grab from the Discovery documentary. It claimed that the colored peaks on the computer screen proved that Tutankhamun belonged to an ancestral line, or haplogroup, called R1b1a2, that is rare in modern Egypt but common in western Europeans.
This immediately led to assertions by neo-Nazi groups that King Tutankhamun had been “white,” including YouTube videos with titles such as King Tutankhamun’s Aryan DNA Results, while others angrily condemned the entire claim as a racist hoax. It played, once again, into the long-running battle over the king’s racial origins. While some worried about a Jewish connection, the argument over whether the king was black or white has inflamed fanatics worldwide. Far-right groups have used blood group data to claim that the ancient Egyptians were in fact Nordic, while others have been desperate to define the pharaohs as black African. A 1970s show of Tutankhamun’s treasures triggered demonstrations arguing that his African heritage was being denied, while the blockbusting 2005 tour was hit by protests in Los Angeles, when demonstrators argued that the reconstruction of the king’s face built from CT scan data was not sufficiently “black.”
For IGENEA, the whole affair was linked to a marketing exercise. It appears to have had no access to the data itself except a snapshot of a computer screen in a TV show, and yet the company now advertises a Tutankhamun DNA Project, which it describes as a search for the pharaoh’s “last living relatives.” The company offers a variety of online DNA tests costing up to $1,500. The sweetener? If your profile matches that of the boy king, you get your money back. Gad refuses to even say whether IGENEA’s analysis of the DNA shown in the documentary is correct. “This is not,” he says, “how science should be conveyed.”
Is there any culture in history that so many are so keen to lay claim to, whether for financial or political gain? “Owning” the pharaohs, it seems, means establishing a privileged place in history to being the founders of civilization. No matter that the ancient Egyptians were almost certainly an ethnically mixed group. They have become a mirror for whoever looks at them, focusing and reflecting the battles and prejudices of today.
Hawass and Gad’s triumphant announcement about Tutankhamun’s family triggered excited media coverage around the world. But what journalists didn’t report was that behind the scenes, the field of ancient DNA was locked in a bitter dispute. A few months later, the Journal of the American Medical Association published a short letter from Eske Willerslev and Eline Lorenzen at the Center for GeoGenetics in Copenhagen, Denmark, one of the world’s most respected ancient DNA labs. It tore Gad’s results and his reputation to shreds.“In most, if not all, ancient Egyptian remains, DNA does not survive to a level that is currently retrievable,” the pair wrote. “We question the reliability of the genetic data presented in this study and therefore the validity of the authors’ conclusions.” Roughly translated, it meant: “we don’t believe a word of it.”
After the heady beginnings of the ancient DNA field in the 1980s and early 1990s, it didn’t take long for the fall. PCR turned out to be extremely susceptible to contamination, far more so than anyone had initially realized. Any trace of modern DNA in the environment a speck of dust, a skin cell, a drop of sweat could dwarf any ancient DNA present and skew the results. In study after study, further analysis revealed that many of the genes that researchers had reported so proudly weren’t ancient at all. Woodward’s 80-million-year-old dinosaur DNA? It actually belonged to a modern human.
Studies of Egyptian mummies were the most controversial of all, because DNA degrades quickly at high temperatures. Although it is possible to retrieve DNA from much older frozen specimens, such as mammoths, the sceptics argued that genetic material from Tutankhamun and his relatives couldn’t possibly have survived 3,000 years in the baking-hot deserts of Egypt. Far from uncovering the secrets of the pharaohs, Gad and his team had been fooled by cross-contamination with modern DNA.
Although Gad and his team wore gloves and masks when working on Tutankhamun, no previous archaeologists had done the same from those unwrapping him in 1925 to those putting him through his CT scan some 80 years later. “You see TV people handling mummies with their bare hands, their sweat dripping on to the mummy,” Tom Gilbert, who heads two research groups at the Center for GeoGenetics, told me. “That’s a classic route of contamination.”
Gad’s team had used other safeguards, including repeating some of their results in a second lab. But critics countered that the team didn’t publish its raw data, and didn’t sequence much of the DNA they amplified. Lorenzen, one of the authors of the letter that attacked Gad’s work, told me: “When working with samples that are so well-known, it is important to convince readers that you have the right data. I am not convinced.” She says she felt obliged to speak out after seeing the huge press coverage the results gained, lamenting that what she saw as flawed conclusions would now be taught in school.
Other prominent scientists shared her concerns. The study “could do a much better job,” complained Svante Pääbo of the Max Planck Institute for Evolutionary Anthropology in Leipzig, Germany, one of the founders of the ancient DNA field. Ian Barnes, an expert in the survival of ancient DNA at the University of London, said he would be “extremely cautious” about using the data. Gilbert, of GeoGenetics, was more blunt. “I’ve given up on the field a long time ago,” he said. “It’s full of crap.”
Gad and his colleagues had been under intense pressure “from Discovery and the forceful Hawass” to get results from incredibly difficult samples. Had they stared into a mix of messy data and contamination and imagined the family relationships they so desperately wanted to see? The team insisted their results were real. They couldn’t prove it, but they were convinced that the elaborate embalming techniques used on Egyptian royalty must have helped to preserve the mummy DNA.
“I don’t understand people’s harshness,” says Carsten Pusch, who joined the Egyptian Museum team soon after the samples were collected. He told me about detailing the months of painstaking experimentation it took to coax DNA from the mummies bones. “These people have never worked with royal mummies. This is pioneering work. I just wish everyone would give us more time.”
 |
 |
 |
 |
 |
We know what a pathetic liar Hawass is; But in fairness, there are other voices.
Ancient Egyptian race controversy
From Wikipedia, the free encyclopedia
When pressed on the issue by American activists in September 2007, the current Secretary General of the Egyptian Supreme Council of Antiquities, Dr. Zahi Hawass stated that "Tutankhamun was not black."
Ahmed Saleh, the former archaeological inspector for the Supreme Council of antiquities stated that the procedures used in the facial re-creation made Tut look Caucasian, "disrespecting the nation's African roots."
In a November 2007 publication of Ancient Egypt Magazine, Hawass asserted that none of the facial reconstructions resemble Tut and that, in his opinion, the most accurate representation of the boy king is the mask from his tomb. The Discovery Channel commissioned a facial reconstruction of Tutankhamun, based on CT scans of a model of his skull, back in 2002.
In molecular evolution, a haplogroup is a group of similar haplotypes that share a common ancestor having the same single nucleotide polymorphism (SNP) mutation in all haplotypes. Haplogroup R-M207 is a Y-chromosome DNA haplogroup. It marks a major split in paleolithic lineages some descendant lines are common throughout Europe, Central Asia and South Asia, and also common in parts of the West Asia and Africa. Others are primarily from West Asia and South Asia. This line is a descendant of haplogroup P-M45.
This haplogroup is believed to have arisen around 20,000-34,000 years ago (Karafet 2008), somewhere in Central Asia or South Asia, where its ancestor Haplogroup P-M45 is most often found at polymorphic frequencies (Wells 2001).
The two currently defined subclades are R-M173 and R-M479. Haplogroup R-M173 is estimated to have arisen during the height of the Last Glacial Maximum (LGM), about 18,500 years ago, most likely in southwestern Asia (Underhill 2009).
Y-haplogroup R-M207 is found throughout all continents, but is fairly common throughout Europe, South Asia and Central Asia. Small frequencies are found in Malaysia, Indonesia, Philippines, and Indigenous Australians (Kayser 2003). It also occurs in Caucasus, Near East, West China, Siberia and some parts of Africa.
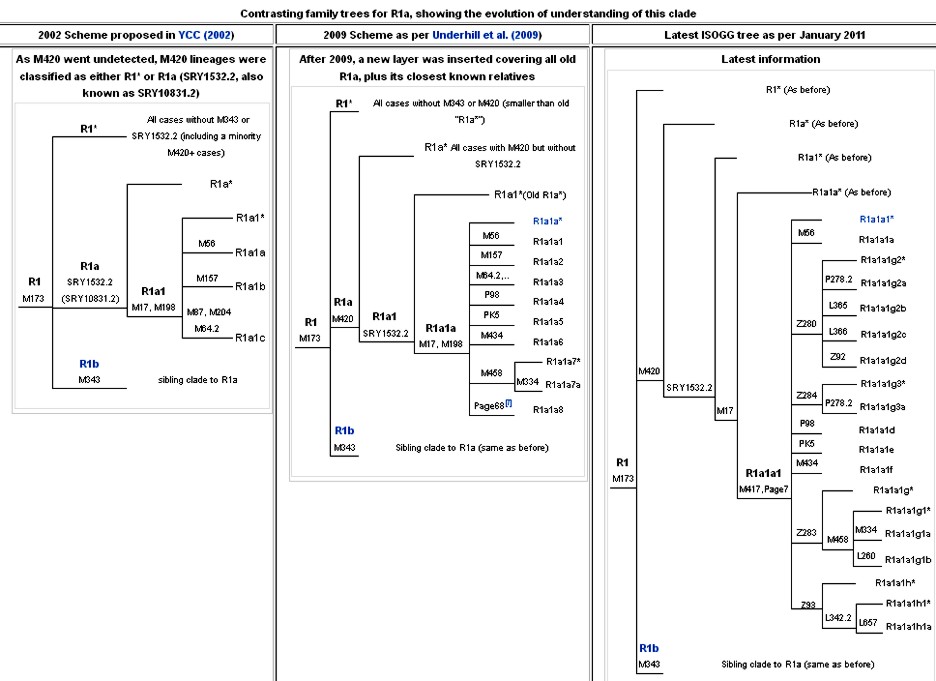 |
R1a and R1a1a are believed to have originated somewhere within Eurasia, most likely in the area from Eastern Europe to South Asia. Several recent studies have proposed that South Asia is the most likely region of origin. But on the other hand, as will be discussed below, some researchers continue to treat modern Indian R1a as being largely due to immigration from the Central Eurasian steppes or Southwestern Asia.
R1a has been found in high frequency at both the eastern and western ends of its core range, for example in India and Tajikistan on the one hand, and Poland on the other. Throughout all of these regions, R1a is dominated by the R1a1a (R-M17 or R-M198) sub-clade.
 |
In South Asia R1a1a has often been observed with high frequency in a number of demographic groups. The main two subclades of R1a1a are R1a1a* and R1a1a7. R1a1a7 is positive for M458 an SNP that separate it from the rest of R1a1a. It is significant because M458 is a European marker and the epicenter is Poland. M458 marker is rare in India.
In India, high percentage of this haplogroup is observed in West Bengal Brahmins (72%) to the east, Konkanastha Brahmins (48%) to the west, Khatris (67%) in north and Iyenger Brahmins (31%) of south. It has also been found in several South Indian Dravidian-speaking Adivasis including the Chenchu (26%) and the Valmikis of Andhra Pradesh and the Kallar of Tamil Nadu suggesting that M17 is widespread in Tribal Southern Indians.
 |
Besides these, studies show high percentages in regionally diverse groups such as Manipuris (50%) to the extreme North East and in Punjab (47%) to the extreme North West.
In Pakistan it is found at 71% among the Mohanna tribe in Sindh province to the south and 46% among the Baltis of Gilgit-Baltistan to the north. While 13% of Sinhalese of Sri Lanka were found to be R1a1a (R-M17) positive.
Hindus of Terai region of Nepal show it at 69%.
In Afghanistan, R1a1a (R-M17) is found at 51.02% among the Pashtuns (the largest ethnic group in Afghanistan) and 30.36% among the Tajiks, but it is less frequent among the Hazaras (6.67%) and the Turkic-speaking Uzbeks (17.65%).
R1a1 among others European haplogrupes
In Europe, R1a, again almost entirely in the R1a1a sub-clade, is found at highest levels among peoples of Eastern European descent (Sorbs, Poles, Russians and Ukrainians; 50 to 65%). In the Baltic countries R1a frequencies decrease from Lithuania (45%) to Estonia (around 30%). Levels in Hungarians have been noted between 20 and 60%.
There is a significant presence in peoples of Scandinavian descent, with highest levels in Norway and Iceland, where between 20 and 30% of men are in R1a1a. Vikings and Normans may have also carried the R1a1a lineage westward; accounting for at least part of the small presence in the British Isles. In East Germany, where Haplogroup R1a reaches a peak frequency in Rostock at a percentage of 31.3%, it averages between 20%-30%.
Haplogroup R1a1a was found at elevated levels amongst a sample of the Israeli population who self-designated themselves as Ashkenazi Jews, possibly reflecting gene flow into Ashkenazi populations from surrounding Eastern European populations, over a course of centuries. This haplogroup finding was apparently consistent with the latest SNP microarray analysis which argued that up to 55 percent of the modern Ashkenazi genome is specifically traceable to Europe. Ashkenazim were found to have a significantly higher frequency of the R-M17 haplogroup Behar reported R-M17 to be the dominant haplogroup in Ashkenazi Levites (52%), although rare in Ashkenazi Cohanim (1.3%) and Israelites (4%).
In Southern Europe R1a1a is not common amongst the general population, but it is widespread in certain areas. Significant levels have been found in pockets, such as in the Pas Valley in Northern Spain, areas of Venice, and Calabria in Italy. The Balkans shows lower frequencies, and significant variation between areas, for example >30% in Slovenia, Croatia and Greek Macedonia, but <10% in Albania, Kosovo and parts of Greece.
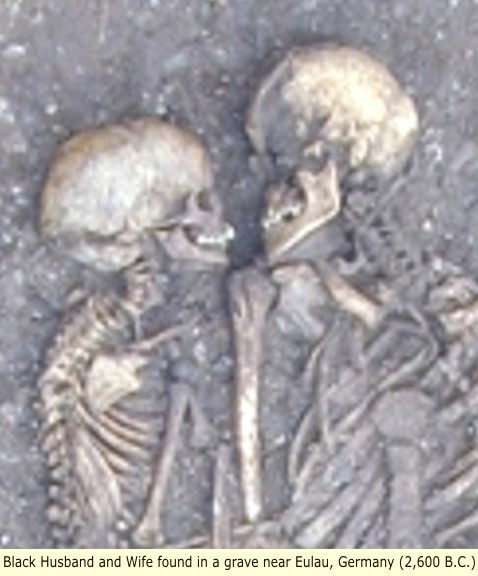
The remains of a father and his two sons, from an archaeological site discovered in 2005 near Eulau (in Saxony-Anhalt, Germany) and dated to about 2600 BCE, tested positive for the Y-SNP marker SRY10831.2. The R1a1 clade was thus present in Europe at least 4600 years ago, in association with one site of the widespread Corded Ware culture.
In 2005 four multiple burials were discovered near Eulau, Germany. The 4,600-year-old graves contained groups of (Black - identified by cranial analysis) adults and children buried facing each other. Skeletal and artifactual evidence and the simultaneous interment of the individuals suggest the supposed families fell victim to a violent event. Genetic analysis of four bodies found in a 4,600-year-old grave shows that they belonged to a mother, a father and their two sons, who were buried together in one another's arms.
 |
 |
R1a1a frequencies are patchy in Central Asia. This variation is possibly a consequence of population bottlenecks in isolated areas and the movements of Scythians in ancient times and later the Turco-Mongols.
High frequencies of R1a1a (R-M17 or R-M198; 50 to 70%) are found among the Ishkashimis, Khujand Tajiks, Panjakent Tajiks, Turkic-speaking Kyrgyzs, and in several peoples of Russia's Altai Republic, but frequencies are relatively lower (16 to 25%) among the Dushanbe Tajiks, Samarkand Tajiks, Yaghnobis and Shughnis.
Although levels are comparatively low amongst some Turkic-speaking groups (e.g. Turks, Azeris, Kazakhs, Yakuts), levels are high (19 to 28%) in certain Turkic or Mongolic-speaking groups of Northwestern China, such as the Bonan, Dongxiang, Salar, and Uyghurs.
In Eastern Siberia, R1a1a is found among certain indigenous ethnic groups including Kamchatkans and Chukotkans, and peaking in Itel'man at 22%.
Middle East and Caucasus
R1a1a has been found in various forms, in most parts of Western Asia, in widely varying concentrations, from almost no presence in areas such as Jordan, to much higher levels in parts of Kuwait, Turkey and Iran.
 |
The Shimar (Shammar) Bedouin tribe in Kuwait show the highest frequency in the Middle East at 43%.
 |
 |
 |
 |
 |
 |
 |
Wells et al. (2001), noted that in the western part of the country, Iranians show low R1a1a levels, while males of eastern parts of Iran carried up to 35% R1a. Nasidze et al. (2004) found R1a in approximately 20% of Iranian males from the cities of Tehran and Isfahan. Regueiro et al. (2006), in a study of Iran, noted much higher frequencies in the south than the north.
Turkey also shows high but unevenly distributed R1a levels amongst some sub-populations. For example Nasidze et al. (2005) found relatively high levels amongst two Kurdish groups of Turkey, the Kurmanji (13%) and Zazaki (26%).
Further to the north of these Middle Eastern regions on the other hand, R1a levels start to increase in the Caucasus, once again in an uneven way. Several populations studied have shown no sign of R1a, while highest levels so far discovered in the region appears to belong to speakers of the Karachay-Balkar language amongst whom about one quarter of men tested so far are in haplogroup R1a1a.
Possible place of origin Southwest Asia
Ancestor R1
Descendants R1b1a (R-P297), R1b1b (R-M335), R1b1c (R-V88)
Defining mutations 1. M343 defines R1b in the broadest sense
P25 defines R1b1, making up most of R1b, and is often used to test for R1b
In some cases, major downstream mutations such as M269 are used to identify R1b, especially in regional or out-of-date studies
Highest frequencies Western Europe, Northern Cameroon, Hazara, Bashkirs
 |
 |
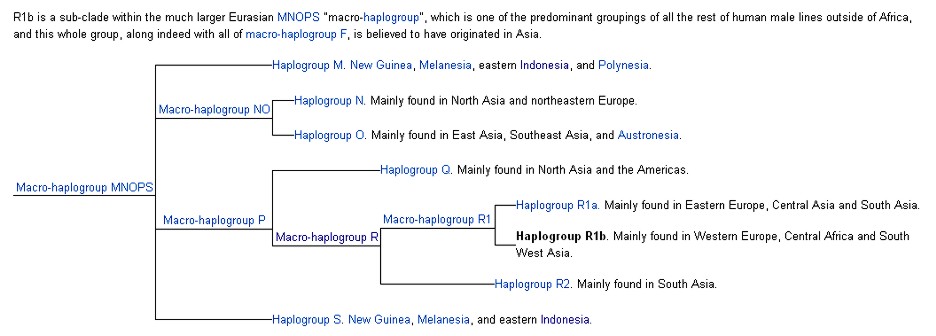
In human genetics, Haplogroup R1b is the most frequently occurring Y-chromosome haplogroup in Western Europe, parts of central Eurasia (for example Bashkortostan), and in parts of sub-Saharan Central Africa (for example around Chad and Cameroon). R1b is also present at lower frequencies throughout Eastern Europe, Western Asia, Central Asia, and parts of South Asia and North Africa.
 |
 |
 |
Due to European emigration it also reaches high frequencies in the Americas and Australia. While Western Europe is dominated by the R1b1a2 (R-M269) branch of R1b, the Chadic-speaking area in Africa is dominated by the branch known as R1b1c (R-V88). These represent two very successful "twigs" on a much bigger "family tree."
| Ireland = 79% Netherlands = 53.5% (Holland) Italy = 49% Germany = 44.5% |
England = 67% France = 61% Belgium = 59.5% Denmark = 44.5% |
 |
 |
R1b1c is found in northern Cameroon in west central Africa at a very high frequency, where it is considered to be caused by a pre-Islamic movement of people from Eurasia.
Suggestive results from other studies which did not test for the full range of new markers discovered by Cruciani et al. have also been reported, which might be in R-V88.
Wood et al. reported high frequencies of men who were P25 positive and M269 negative, amongst the same north Cameroon area where Cruciani et al. reported high R-V88 levels. However they also found such cases amongst 3% (1/32) of Fante from Ghana, 9% (1/11) of Bassa from southern Cameroon, 4% (1/24) of Herero from Namibia, 5% (1/22) of Ambo from Namibia, 4% (4/92) of Egyptians, and 4% (1/28) of Tunisians.
 |
 |
 |
Luis et al. found the following cases of men M173 positive (R1), but negative for M73 (R1b1b1), M269 (R1b1b2), M18 (R1b1a1, a clade with V88, M18 having been discovered before V88) and M17 (R1a1a): 1 of 121 Omanis, 3 of 147 Egyptians, 2 of 14 Bantu from southern Cameroon, and 1 of 69 Hutu from Rwanda.
Pereira et al.
(2010) In a study of several Saharan Tuareg populations, found one third of 31 men tested from near Tanut in Niger to be in R1b.
 |
 |
 |
 |
Historical note
The DNA tests that assisted in the identification of Czar Nicholas II of Russia found that he had haplogroup R1b.
 |
R1b1c1 (R-M18)
R1b1c1 is a sub-clade of R-V88 which is defined by the presence of SNP marker M18. It has been found only at low frequencies in samples from Sardinia and Lebanon.
Haplogroup R-M124 is a Y-chromosome haplogroup characterized by genetic markers M124, P249, P267, L266, and is mainly found in South Asia, parts of Central and West Asia.
Haplogroup R-M124, along with haplogroups H, L, R1a1, and J2, forms the majority of the South Asian male population. The frequency is around 10-15% in India and Sri Lanka and 7-8% in Pakistan. Its spread within South Asia is very extensive, ranging from Baluchistan in the west to Bengal in the east; Hunza in the north to Sri Lanka in the south.
North Indian Muslims have a frequency of 11%(Sunni) and 9%(Shia), while Dawoodi Bohra Muslim in the western state of Gujarat have a frequency of 16% and Mappla Muslims of South India have a frequency of 5%. The R-M124 haplogroup is also found in 14% of the Burusho people who speak the language isolate called Burushaski.
Some of the other studies like Bamshad et al., 2001, Kivisild et al., 2003 found Haplogroup 1 (the old representation for non-R1a1 Haplogroup R subclades) at around 40% among Telugus of coastal Andhra Pradesh. The identification of this Haplogroup with R-M124 is confirmed from Sanghamitra Sahoo et al., 2006 study which observed R-M124 ranging from 35% to 55% among non-Brahmin castes of this region.
Haplogroup R-M124 comprises 53% of Y-chromosomes among Sinti, a subgroup of the Romani people living in Germany who were relocated to Central Asia, however the sample size was only 15 individuals. This Romani branch has its ancient roots in India.
Central Asia
In Central Asia, Tajikistan shows Haplogroup R-M124 at 6%, while the other '-stan' states vary around 2%. Bartangis of Tajikistan have a high frequency of R-M124 at about 17%, Ishkashimi at 8%, Khojant at 9% and Dushanbe at 6%.
Specifically, Haplogroup R-M124 has been found in approximately 7.5% (4/53) of recent Iranian emigrants living in Samarkand,[10] 7.1% (7/99) of Pamiris,[10] 6.8% (3/44) of Karakalpaks, 5.1% (4/78) of Tajiks, 5% (2/40) of Dungans in Kyrgyzstan, 3.3% (1/30) of Turkmens, 2.2% (8/366) of Uzbeks, and 1.9% (1/54) of Kazakhs.
West Asia
One study has found Haplogroup R-M124 at an unusually high frequency of 44% (11/25) among Kurmanji speakers (Kurmanjs) in Georgia, but at a much lower frequency of 8% (7/87) among Kurmanjs in Turkey.
An R-M124 frequency of 15.8% was observed among Chechens. R-M124 has been found in approximately 8% (2/24) of a sample of Ossetians from Alagir.
In the Caucasus, around 16% of Mountain Jews, 8% of Balkarians,[14] 6% of Kalmyks,[15] 3% of Azerbaijanis,[12] 2.6% of Kumyks,[16] 2.4% of Avars,[16] 2% of Armenians, and 1% to 6% of Georgians belong to the R-M124 haplogroup. Approximately 1% of Turks and 1% to 3% of Iranians also belong to this haplogroup.
Arab World
In the R2-M124-WTY and R-Arabia Y-DNA Projects, Haplogroup R-M124 has appeared in the following Arab countries: Kuwait (3 clusters), Saudi Arabia (2 clusters), United Arab Emirates (1 cluster), Syrian Arab Republic (1 cluster), and Tunisia (1 cluster).
Thus, Haplogroup R-M124 has been observed among Arabs at low frequencies in 11 countries/territories (Egypt, Jordan, Kuwait, Lebanon, Palestine, Qatar, Saudi Arabia, Syria, Tunisia, United Arab Emirates, and Yemen) of the 22 Arab countries/territories so far.
HAPLOGROUP R
Haplogroup R, the ancestral clade to R1 and R2, appeared on the Central Asian Steppes around 35,000 to 30,000 years ago.
R1, sister clade to R2, moved to the West from the Central Asian Steppes around 35,000 to 30,000 years ago. R1 pockets were established, from where R1a and R1b emerged.
R2a [R-M124] made its first entry into the Indian sub-continent around 25,000 years ago. The routes taken are not clear, although the Indus and Ganges rivers are possible theories put forward. There could, of course, have been multiple immigrations of this haplogroup into the Indian sub-continent, both in the Paleolithic and the Neolithic.
At least 90% of R-M124 individuals are located in the Indian sub-continent. It is also reported in Caucasus and Central Asia.
Haplogroup R2a is present both in Dravidian and other Indian populations, meaning that R2a has a pan-Indian presence, and not restricted to any linguistic group.
Haplogroup R2a has a more significant presence in middle and upper castes. The frequencies of R2a seem to mirror the frequencies of R1a (i.e. both lineages are strong and weak in the same social and linguistic subgroups). This may indicate that both R1a and R2a moved into India at roughly the same time or cohabited, although more research is needed.
R1a1 and R2a haplogroups indicate demographic complexity that is inconsistent with a recent single history and is not inconsistent with a more proximal Central Asian input of the R2a haplogroup in the upper castes. R2a has a particularly strong presence in the Indian states of West Bengal, Uttar Pradesh and Gujarat, and in the area of Mumbai (Bombay).
The paper claims that there is no evidence that Central Asia was the source of the R1a and R2a lineages in India. The theory that Central Asia could have been the recipient of the two lineages from India should not be ruled out. (Comment: The Dravidian Albinos moving to Central Asia from India accounts for this). In addition, the data are not inconsistent with complex exchanges of this haplogroup between Central Asia and the Indian sub-continent, with the latter being both the source and the recipient at different times.
Haplogroup K-M526 (Y-DNA).
K(xLT) is the ancestral haplogroup to haplogroups K1, K2, K3, K4, M, NO, P (which contains haplogroups Q and R), and S (formerly MNOPS). Possible time of origin 35,000-45,000 years BP in South or Central Asia.
Haplogroup P-M45 (Y-DNA) is the parent of haplogroups (P*, Q, R). It is believed to have arisen 27,000-41,000 years BP in Central Asia - South Asia.
This haplogroup contains the patrilineal ancestors of most Europeans and almost all of the indigenous peoples of the Americas. It also contains approximately one third to two thirds of the males among various populations of Central Asia and Southern Asia.
Haplogroup R-M207 (Y-DNA)
In human population genetics, haplogroups define the major lineages of direct paternal (male) lines back to a shared common ancestor in Africa.
haplogroup R-M207 is a Y-chromosome DNA haplogroup. It marks a major split in paleolithic lineages some descendant lines are common throughout Europe, Central Asia and South Asia, and also common in parts of the West Asia and Africa. Others are primarily from West Asia and South Asia. This line is a descendant of haplogroup P-M45.
This haplogroup is believed to have arisen around 20,000-34,000 years ago,(Karafet 2008) somewhere in Central Asia or South Asia, where its ancestor Haplogroup P-M45 is most often found at polymorphic frequencies.(Wells 2001)
The two currently defined subclades are R-M173 and R-M479. Haplogroup R-M173 is estimated to have arisen during the height of the Last Glacial Maximum (LGM), about 18,500 years ago, most likely in southwestern Asia.
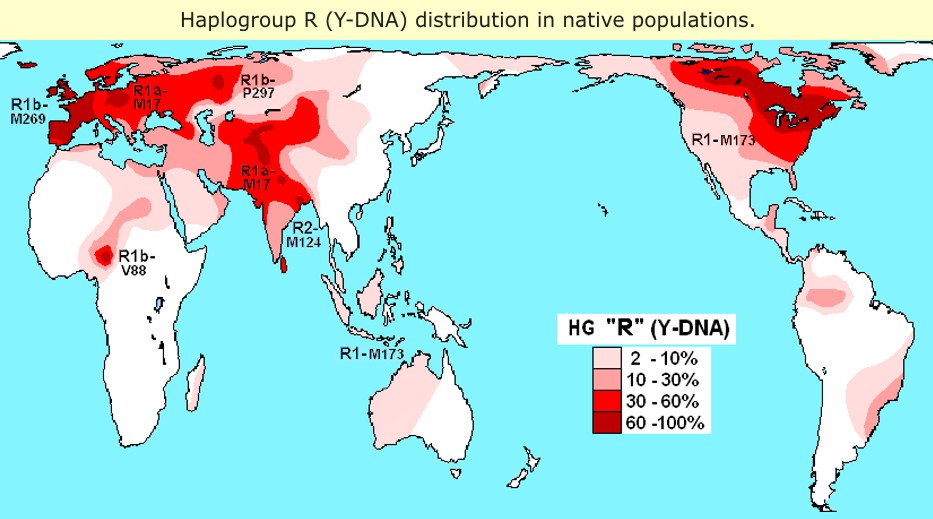
Y-haplogroup R-M207 is found throughout all continents, but is fairly common throughout Europe, South Asia and Central Asia. Small frequencies are found in Malaysia, Indonesia, Philippines, and Indigenous Australians.(Kayser 2003) It also occurs in Caucasus, Near East, West China, Siberia and some parts of Africa. It has a high frequency in the Native Americans due primarily to the introduction of Eurasian lineages in the last 500 years.
Haplogroup R-M173 (Y-DNA)
In human genetics, Haplogroup R-M173 is a Y-chromosome DNA haplogroup, a subgroup of haplogroup R, associated with the M173 mutation. It is dominated in modern populations by two Eurasian clades, R-M240 and R-M343, which together are found all over Eurasia except in Southeast Asia and East Asia. However, other types of R-M173, less well-known and undefined so far by any identified SNP, and therefore referred to collectively simply as R-M173*, have been reported in the Americas, all over Asia and Oceania.
In the Americas, it is not a pre-Colombian founding lineage. However, it is the second most common haplogroup in Indigenous peoples of the Americas following haplogroup Q-M242, and spreads specially in Algonquian peoples from United States and Canada.
The origins of R-M173 remain unclear. Haplogroup R-M207 is part of the family of haplogroup P-M45, and a sibling clade, therefore, of haplogroup Q-M242, which is common in the Americas and Eurasia. In Eurasia, Q-M242's geography includes eastern areas such as Siberia. Based on these ancestral lineages, an inferred origin for R-M173 to the east of the West Asia. For example, Kivisild 2003 believes the evidence "suggests that southern and western Asia might be the source of this haplogroup." and "Given the geographic spread and STR diversities of sister clades R1 and R2, the latter of which is restricted to India, Pakistan, Iran, and southern central Asia, it is possible that southern and western Asia were the source for R1 and R1a differentiation." Soares 2010 felt in their review of the literature, that the case for South Asian origins is strongest, with the Central Asian origin argued by (Wells 2001) being also worthy of consideration.
Haplogroup R-M173 is fairly common throughout Europe, South Asia and Central Asia. It also occurs in Africa, Near East and Native Americans from North America. Low frequencies in Siberia, Malay Archipelago and Indigenous Australians.
In Indigenous Americans groups, R-M173 is the most common haplogroup after the various Q-M242, especially in North America in Ojibwe people at 79%, Chipewyan 62%, Seminole 50%, Cherokee 47%, Dogrib 40% and Papago 38%. The decreasing gradient of haplogroup R-M207 from Northeastern to Southwestern North America is evidence that this results from European admixture.
 |
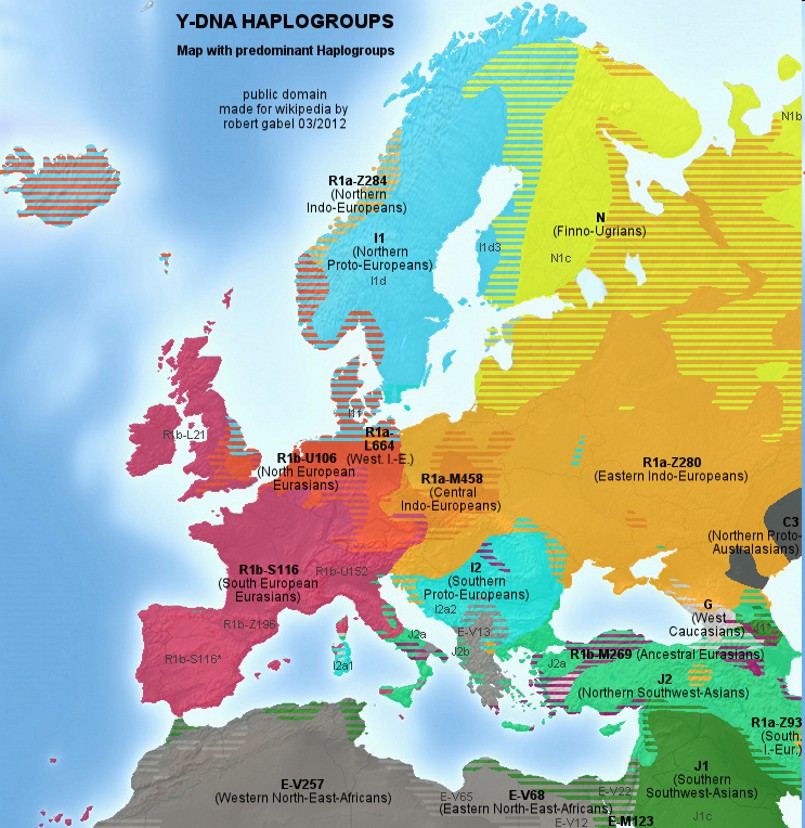 |
Haplogroup I-M170 is a Y-chromosome DNA haplogroup, a subgroup of haplogroup IJ, itself a derivative of Haplogroup IJK. Y-DNA Haplogroup I-M170 is now predominantly a European haplogroup. It can be found in the majority of present-day European populations with peaks in some Northern and South-Eastern European countries where the total population is small in comparison with European standards. Consequently, the haplogroup represents not more than one-fifth of the population of Europe, being the continent's second major Y-DNA haplogroup behind Haplogroup R. Haplogroup I-M170 Y-chromosomes have also been found among some populations of the Near East, the Caucasus, Northeast Africa and Central Siberia. The haplogroup reaches its maximum frequency in the Dinaric Alps (with the highest concentration in present-day Bosnia and Herzegovina).
 |
Haplogroup N (M231) is a Y-chromosome DNA haplogroup typical of northern Eurasia, defined by the presence of the marker M231. Haplogroup N-M231 is a descendant haplogroup of Haplogroup NO. It is considered relatively young, having populated the north of Eurasia after the last Ice Age. Males carrying the marker apparently moved northwards as the climate warmed in the Holocene.
Haplogroup N-231 has a wide geographic distribution throughout northern Eurasia, and it also has been observed occasionally in more southerly areas, including Southeast Asia, Nepal, Southwest Asia, and Southern Europe. Its highest frequency occurs among the Finnic and Baltic peoples of northern Europe, the Ob-Ugric and Northern Samoyedic peoples of western Siberia, and the Siberian Turkic-speaking Yakuts (McDonald 2005).
 |
OCA1 is caused by an alteration of the tyrosinase gene, and can occur in two variations. The first is OCA1a, and means that the organism cannot develop pigment at all. The hair is usually white (often translucent) and the skin very pale. Vision usually ranges from 20/200 to 20/400. The second is OCA1b, which has several subtypes itself. Some individuals with OCA1b can tan and also develop pigment in the hair. One subtype of OCA1b is called OCA1b TS (temperature sensitive), where the tyrosinase can only function below a certain temperature, which causes the body hair in cooler body regions to develop pigment (i.e. get darker). (An equivalent mutation produces the coat pattern in Siamese cats. Another variant of OCA1b, called Albinism, yellow mutant type (OMIM: 606952) is more common among the Amish than in other populations, and results in blonde hair and the eventual development of skin pigmentation during infancy, though at birth is difficult to distinguish from other types.
 |
 |
The most common type of albinism, is caused by mutation of the P gene. People with OCA2 generally have more pigment and better vision than those with OCA1, but cannot tan like some with OCA1b. A little pigment can develop in freckles or moles. People with OCA2 usually have fair skin but often not as pale as OCA1, and pale blonde to golden, strawberry blonde, or even brown hair, and most commonly blue eyes.
 |
 |
 |
|
| These three sisters all have the EXACT same haplogroup. The only difference is that the one in the middle has a damaged "P" gene. |
These two brothers have the EXACT same haplogroup. The only difference is that the one on the right has a damaged "P" gene. |
 |
| This brother and his two sisters all have the EXACT same haplogroup. The only difference is that the one in the middle has a Undamaged "P" gene. |
 |
 |
Eupedia.comDistribution of European Y-chromosome DNA (Y-DNA) haplogroups by region in percentageLast update : February 2010 (Armenians, Azeris, Basques, Bashkirs, Bosnians, Cantabrians, Cypriots, Galicians, Kurds, Macedonians) Human Y-chromosome DNA can be divided in genealogical groups sharing a common ancestor. These are called haplogroups . To know what ancient ethnic group is associated with each haplogroup, please check European Haplogroups : origins, geographic spread and relation to ethnic groups . Note that figures are only indicative. Several sources were used and averages recalculated by merging the data available. Being approximations, numbers were rounded up to 0.5%. Frequencies inferior to 0.25% are indicated as 0%. A non-exhaustive list of the sources used for this page can be found here . Note: the number in each ROW indicates a percentage of the population, relative to each haplogroup. |
 |
 |
NotesTurkey is the only country that includes a sizeable percentage of Asian and African haplogroups not listed in this table (A, ExE1b1b, C, H, L, O, R2) representing 8.5% of the total. Haplogroup L alone makes up 4% of the Turkish population.The division of Italy is as follows: North Italy is everything until Liguria and Emilia-Romagna; Central Italy comprises Tuscany, Marche, Umbria, Latium and Abruzzo. South Italy is everything else to the south, except Sardinia and Sicily, which have been made into separate categories due to their specific history and relative geographic isolation. Sources for the Italian regional breakdown . Our division of Germany was made this way : North Germany includes the Schleswig-Holstein, Lower Saxony (+ Hamburg and Bremen) and Mecklenburg-Western Pomerania. West Germany is the Rhineland, Hesse and Saarland. South Germany is Baden-Württemberg and Bavaria. East Germany is composed of Brandenburg, Berlin, Saxony-Anhalt, Saxony and Thuringia. The sample size for each country or region is at least of 100. Italy, Germany, England and Ireland have over 2000 samples each, France and Spain over 1000, Portugal over 900, Belgium over 750, the Netherlands, Finland and Hungary over 650, Greece and Turkey over 500. Surrounding regions |
 |
 |
 |
Additional information
The percentages of haplogroups H1, H3 and U5 is given in addition to the total for H and U. This is useful to assess the proportion of Paleolithic European (Cro-Magnon) lineages, as opposed to later arrivals.
The "Other" category includes mostly the older haplogroups N, R, pre-HV and HV, but also occasionally a few African (L) or Asian haplogroups (A, B, C, D, M, Z).
The largest sample sizes in this data base are Germany (n = 2610), England (n = 1577), Scotland (n = 1413), Ireland (n = 1397), France (n = 878), Italy (n = 808), Norway (n = 703), Finland (n = 580), and Iceland (n = 511). Each country has at least 100 samples.
 |
 |
 |
From Dienekes Pontikos
http://dienekes.blogspot.com/
(Last Updated 12 Oct 2013) .
Neolithic Linearbandkeramik from Derenburg [2 F*(xG,H,I,J,K), 1 G2a3]
Neolithic Spain [5 G2a, 1 E-V13]
Hongshan culture Neolithic China [1 C, 1 O3, 4 N1(?N1a,N1c)]
Longshan culture Neolithic China [3 N1(xN1a,N1c)]
Xiaoheyan culture Neolithic China [12 N1(xN1a,N1c)]
Neolithic Ötzi from the Alps [G2a4]
Prehistoric Alaskan [Q3]
Prehistoric South Siberians from Krasnoyarsk and here [10 R1a1, 1 C(xC3)]
Neolithic southwestern France from Treilles [20 G2a, 2 I2a]
Neolithic Megalithic France from la Pierre Fritte [2 I2a1]
Neolithic Bell Beaker from Kromsdorf Germany [2 R1b]
Bronze Age from West Liao-River northern China [N-M231, O3-M122]
Lower Xiajiadian Bronze Age West Liao-River northern China [3 N1(xN1a,N1c, 2 O3]
Upper Xiajiadian Bronze Age West Liao-River northern China [1 C3e, 3 N1c, 1 N1(xN1a,N1c), 2 O3a, 2 O3a3c]
Northern Steppe culture Bronze Age West Liao-River northern China [12 C3e]
Bronze Age from Tarim basin in Xiaohe [7 R1a1a]
Prehistoric Paleo-Eskimo from Greenland [1 Q1a]
Ancient Chinese from the Yangtze River [14 O1, 3 O2a, 7 O3*, 5 O3d, 1 O3e, 18 undetermined]
Ancient nomads from Pengyang China [4 Q]
Eneolithic Corded Ware Germans [3 related R1a]
Bronze Age Lichtenstein Cave in Germany [estimated presence I1b2*, R1a1, R1b1c]
Ancient Mongolian [presence of Tat-C in Yakut and Xiongnu]
Ancient Egyin Gol Mongolians and here and here [Y-STR in Table 2 of second study; N3, Q, C]
Ancient Mongolian Xiongnu [1 R1a1]
New Kingdom Egyptian pharaoh Ramesses III [1 E1b1a]
Aboriginals from Canary Islands [E-M78, E-M81, J-M267, E-M33, I-M170, K-M9, P-M45, R-M269]
Late Antique Basques [4 I, 2 R1b3d, 19 R1(xR1a1), 2 R-M173]
Late Antique Imperial Roman from Bavaria [2 R1b, 2 I1, 2 E1b1b, 2 I1/G2a]
Medieval Hungarians [Two Tat-C out of four]
Medieval Germans from Ergolding, Bavaria, Germany [4 R1b (two siblings), 2 G2a]
Medieval Germans (?) from Usedom, Mecklenburg-Vorpommern, Germany [E1b1b, R1a1a7]
Medieval Swedes from Stockholm [2 I1, probably related]
Recent Frozen Yakuts [8 N1c, 5 non-N1c]
Human remains excavated in a Spanish funeral cave dating from the beginning of the fifth millennium B.C. [G2a and E1b1b1a1b].
A European population in Minoan Bronze Age Crete [The Minoan mtDNA haplotypes resembled those of the European populations. The majority of Minoans were classified in haplogroups H (43.2%), T (18.9%), K (16.2%) and I (8.1%). Haplogroups U5A, W, J2, U, X and J were each identified in a single individual]
Look-up mtDNA Haplogroup H at Wiki:
In human mitochondrial genetics, Haplogroup H is a human mitochondrial DNA (mtDNA) haplogroup that likely originated in Southwest Asia 20,000-25,000 YBP.
Haplogroup H is a descendant of haplogroup HV. The Cambridge Reference Sequence (CRS), the human mitochondrial sequence to which all other sequences are compared, belongs to haplogroup H2a2a. Several independent studies conclude that haplogroup H probably evolved in West Asia c. 25,000 years ago. It was carried to Europe by migrations c. 20-25,000 years ago, and spread with population of the southwest of the continent. Its arrival was roughly contemporary with the rise of the Gravettian culture. The spread of subclades H1, H3 and the sister haplogroup V reflect a second intra-European expansion from the Franco-Cantabrian region after the last glacial maximum, c. 13,000 years ago. However, any statements concerning the geographic origin of this or any other haplogroup are highly speculative and considered by most population geneticists to be 'story telling' and outside the domain of science. Furthermore, inferring close associations between a haplogroup and a specific archaeological culture can be equally problematic.
In July 2008 ancient mtDNA from an individual called Paglicci 23, whose remains were dated to 25,000 years ago and excavated from Paglicci Cave (Apulia, Italy), were found to be identical to the Cambridge Reference Sequence in HVR1. This once was believed to indicate haplogroup H, but researchers now recognize that CRS can also appear in U or HV.
Haplogroup H is the most common mtDNA haplogroup in Europe. Haplogroup H is found in approximately 41% of native Europeans. The haplogroup is also common in North Africa and the Middle East. The majority of the European populations have an overall haplogroup H frequency of 40%–50%. Frequencies decrease in the southeast of the continent, reaching 20% in the Near East and Caucasus, 17% in Iran, and <10% in the Persian Gulf, Northern India and Central Asia.
Subhaplogroups H1 and H3
Among all these clades, the subhaplogroups H1 and H3 have been subject to a more detailed study and would be associated to the Magdalenian expansion from SW Europe c. 13,000 years ago:
H1 encompasses an important fraction of Western European mtDNA, reaching its local peak among contemporary Basques 27.8% and appearing at a high frequency among other Iberians and North Africans. Its frequency is above 10% in many other parts of Europe (France, Sardinia, British Isles, Alps, large portions of Eastern Europe), and above 5% in nearly all the continent. Its subclade H1b is most common in eastern Europe and NW Siberia. So far, the highest frequency of H1 - 61%- has been found among the Tuareg of the Fezzan region in Libya.
| Click for Realhistoryww Home Page |