It is my conviction that a white skin is not natural to man, and that by nature he has either a black or brown skin like our forefathers, the Hindoos, and that the white man was never originally created by nature; and that, therefore, there is no race of white people. From...METAPHYSICS OF SEXUAL LOVE by Arthur Schopenhauer.
The University of Adelaide Library (pdf)
|
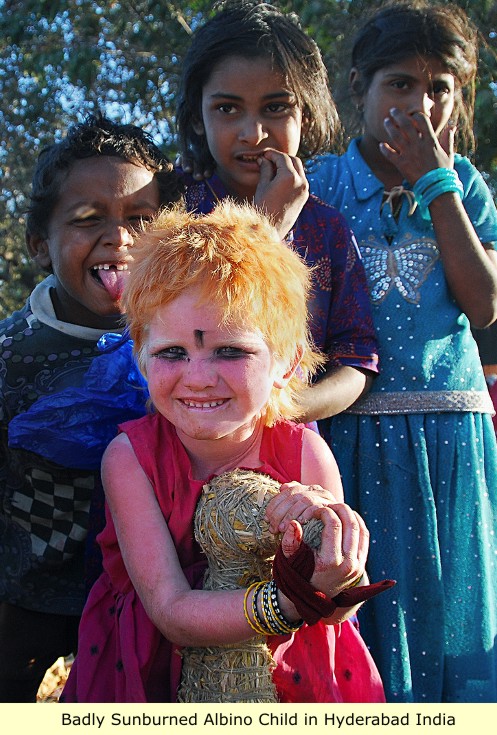
The subject matter of Albinism has come to require so much attention because Europeans are the Albinos of India's indigenous Dravidians, who split from India and went into Central Asia many thousands of years ago. But Europeans steadfastly refuse to admit this, even though this obvious conclusion is supported by simple observation - They look exactly alike - except for skin color. And modern "Dravidian Albinos" are indistinguishable from Europeans. Combine that with the genetic data, and there is no doubt that they are the same people.
 |
 |
 |
 |
 |
 |
 |
|
Haplogroup R (Y-DNA)
|
Background on Haplogroup "R"
Origins of Y-dna Haplogroup "R"
From Wikipedia: According to the Genographic Project conducted by the National Geographic Society, Haplogroup R2a arose about 25,000 years ago in Central Asia and its members migrated southward as part of the second major wave of human migration into India. According to Sengupta et al. (2006), uncertainty neutralizes previous conclusions that the intrusion of HGs R1a1 and R2 [Now R-M124] from the northwest in Dravidian-speaking southern tribes is attributable to a single recent event. Rather, these HGs contain considerable demographic complexity, as implied by their high haplotype diversity. Specifically, they could have actually arrived in southern India from a southwestern Asian source region multiple times, with some episodes considerably earlier than others. The following is Manoukian's (2006) summary of the findings of the Genographic Project conducted by the National Geographic Society and directed by Spencer Wells (2001): Haplogroup R, the ancestral clade to R1 and R2, appeared on the Central Asian Steppes around 35,000 to 30,000 years ago. R1, sister clade to R2, moved to the West (READ EUROPE) from the Central Asian Steppes around 35,000 to 30,000 years ago. R1 pockets were established, from where R1a and R1b emerged. R2a [R-M124] made its first entry into the Indian sub-continent around 25,000 years ago. The routes taken are not clear, although the Indus and Ganges rivers are possible theories put forward. There could, of course, have been multiple immigrations of this haplogroup into the Indian sub-continent, both in the Paleolithic and the Neolithic |
The final proof that Europeans are Albinos derived from Dravidian Indians, is the Genetic Distance Maps created by the studies:"Genetic Distance Map from The History and Geography of Human Genes" by Cavalli-Sforza.And "The Genetic Structure and History of Africans and African Americans" by Sarah A. Tishkoff.Both genetic maps show that Black and Brown Indians, and White Europeans, are alone together, separate from all other humans, like two peas in a pod. The only difference is that one group is pigmented, and the other is not! One group is Albino, the other is not!
|
 |
But of course though Europeans are mostly derived, they are not solely derived, from Dravidian Albinos, or haplogroup "R":
European Albinos are also in substantial part, the Albinos of Africans carrying haplogroups I, E, and J, among others.
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
In order to understand Albinism, the genetic machinations which cause Albinism, and the scientific terms used to describe these machinations, must be understood.
In all the cells of our body, our genes are found on chromosomes (long strings of genes). We have many thousands of genes that provide information for our body to grow, develop and remain healthy. The gene sends messages to the cell to make important chemical products such as proteins.
There are usually 46 chromosomes in each cell that are arranged into 23 pairs. One of each pair is passed on to us from our mother and the other from our father. 22 of these chromosome pairs are numbered. These numbered pairs are known as the autosomal chromosomes. The 23rd pair is made up of the sex chromosomes called X and Y. Males have an X and a Y chromosome and females have two copies of the X chromosome.
Since the chromosomes come in pairs, there are also two copies of each of the genes. The exception to this rule applies to the genes carried on the sex chromosomes called X and Y. A variation in a gene that creates a fault is called a pathogenic variant or mutation. Genes are sections of DNA that code for the proteins our body needs to function. A mutation in a gene will affect the body differently depending on how much it changes the resulting protein, how critical that protein is to the body and how much of that protein is needed in the body.
If a DNA change occurs in only one of the pair of genes and this causes a health condition, it is called a dominant mutation. If a health condition only occurs when both copies of the gene are changed, this is called a recessive mutation.
Humans contain two copies of each gene, one from the father and one from the mother, which sometimes are referred to as the alleles of a gene. If a mutation occurs in just one copy of the gene then that individual is considered heterozygous. On the other hand if both copies of a gene are mutated then that individual is homozygous genotype. Majority of hereditary disorders are harmful if both copies or alleles of a gene are affected, which means protein products from both genes may fail to operate properly. In such cases immediate medical attention is needed so the function of a defected protein can be restored through medication. In heterozygous genotypes one copy of the gene is healthy and can produce fine proteins thus these individuals are usually not affected and are considered just carriers. However in a few hereditary disorders heterozygous individuals may suffer from a milder version of the disease.
An autosome is a chromosome that is not an allosome (a sex chromosome). Autosomes appear in pairs whose members have the same form but differ from other pairs in a diploid cell, whereas members of an allosome pair may differ from one another and thereby determine sex.
Oculocutaneous albinism is inherited in an autosomal recessive pattern, which means both copies of a gene in each cell have mutations. Most often, the parents of an individual with an autosomal recessive condition each carry one copy of the mutated gene, but they do not show signs and symptoms of the condition.
Autosomal recessive is one of several ways that a trait, disorder, or disease can be passed down through families. An autosomal recessive disorder means two copies of an abnormal gene must be present in order for the disease or trait to develop.
There are several different types of Albinism which affect several different genes. Each is "exclusive" and causes a DIFFERENT type of Albinism.
Example:
If two people with the same type of albinism reproduce (such as two OCA-1 people), all of their children will have OCA-1 albinism. If two people with two different types of albinism have children (such as one parent being OCA-1 and the other parent being OCA-2): In the case of Blacks - NONE of their children will have albinism, because no PAIR of the SAME gene is created: (meaning that the children will look the same as their "Unaffected" parents would look. In the case of Europeans, because they have the fixed phenotype of a OCA-2, the children will look like a standard European. i.e. Since Europeans are already OCA-2 the child will look OCA-2
It seems that as Blacks have gained access to the information about Albinism, and logically/truthfully interpreted it. The Albinos realized the threat the truth posed to them: (what with all their lies about themselves and their history). So they have been busy over the last few years, re-writing material on Albinism, creating new lies, and just generally muddying the water, to make it harder to understand Albinism.
Study: Molecular basis of albinism: Mutations and polymorphisms of pigmentation genes associated with albinism
First published: 6 April 1999 -
Mutations in six genes have been reported to be responsible for different types of oculocutaneous and ocular albinism, including the tyrosinase gene (TYR) and OCA1 (MIM# 203100), the OCA2 gene and OCA2 (MIM# 203200), the tyrosinase-related protein-1 gene (TYRP1) and OCA3 (MIM# 203290), the HPS gene and Hermansky-Pudlak syndrome (MIM# 203300), the CHS gene (CHS1), and Chediak-Higashi syndrome (MIM# 214500), and the X-linked ocular albinism gene and OA1
Dr. Orville Boyd Jenkins
The origins of Europeans have come into sharp focus in the past year as researchers have sequenced the genomes of ancient populations, rather than only a few individuals. By comparing key parts of the DNA across the genomes of 83 ancient individuals from archaeological sites throughout Europe, the international team of researchers reported earlier this year that Europeans today are a mix of the blending of at least three ancient populations of hunter-gatherers and farmers who moved into Europe in separate migrations over the past 8000 years. The study revealed that a massive migration of Yamnaya herders from the steppes north of the Black Sea may have brought Indo-European languages to Europe about 4500 years ago.
When it comes to skin color, the team found a patchwork of evolution in different places, and three separate genes that produce light skin, telling a complex story for how European’s skin evolved to be much lighter during the past 8000 years. The modern humans who came out of Africa to originally settle Europe about 40,000 years are presumed to have had dark skin. which is advantageous in sunny latitudes. The new data confirm that about 8500 years ago, early hunter-gatherers in Spain, Luxembourg, and Hungary also had darker skin: They did not have mutations in two genes—SLC24A5 and SLC45A2—that lead to depigmentation and, therefore, pale skin (Albinism) in Europeans today. [Edited to remove some Albino bull].
WHITE SKIN GENESSince the discovery of "Skin Color" genes: Albino scientists have been going all over Europe trying to find ancient Skeletons in Europe with these genes, in order to prove that Whites were in Europe before circa 1,200 B.C. Of course they didn't explain to White people just what those genes were about, and how they indicated when a Skeleton was of a White person or a Black person. See below. |
 |
Scientific Study: Derived Immune and Ancestral Pigmentation Alleles in a 7,000-Year-old Mesolithic European (2014)
Quote: For each variant we assessed whether the Mesolithic genome carried the ancestral or derived (putatively adaptive) allele. Of the ten variants, the Mesolithic genome carried the ancestral and non-selected allele as a homozygote in three regions: C12orf29 (a gene with unknown function), SLC45A2 (rs16891982) and SLC24A5 (rs1426654) (Table 1). The latter two variants are the two strongest known loci affecting light skin pigmentation in Europeans and their ancestral alleles and associated haplotypes are either absent or segregate at very low frequencies in extant Europeans (3% and 0% for SLC45A2 and SLC24A5 respectively) (Fig. 2). We subsequently examined all genes known to be associated with pigmentation in Europeans, finding ancestral alleles in MC1R, TYR and KITLG and derived alleles in TYRP1, ASIP and IRF4 (Supplementary Information). Although the precise phenotypic effects cannot currently be ascertained in a European genetic background, results from functional experiments indicate that the allelic combination in this Mesolithic individual is likely to have resulted in dark skin pigmentation and dark/brown hair. Further examination revealed that this individual carried the rs12913832*C single nucleotide polymorphism (SNP) and the associated homozygous haplotype spanning the HERC2/OCA2 locus that is strongly associated with blue eye color. Moreover, a prediction of eye color based on genotypes at additional loci using HIrisPlex24 yielded a 0.823 maximal and 0.672 minimal probability for being non-brown eyed (Supplementary Information). The genotypic combination leading to a predicted phenotype of dark skin and non-brown eyes is unique and no longer present in contemporary European populations. Our results indicate that the adaptive spread of light skin pigmentation alleles was not complete in some European populations by the Mesolithic, and that the spread of alleles associated with light/blue eye color may have preceded changes in skin pigmentation.
Derived means = to obtain something from (a specified source) (a direct connection).
Putatively means = generally considered or reputed to be the case (a guess). These lying Albinos started off lying!
Ancestral gene means = DNA sequences in animal species that lived millions of years ago but that, although much modified by mutation and rearrangement are still present in whole or in part in the contemporary human genome. Some distinct and separate human genes share sequences that suggest they may have had a common ancestry.
Explanation of the OMIM column below: Online Mendelian Inheritance in Man (OMIM) is a continuously updated catalog of human genes and genetic disorders and traits, with a particular focus on the gene-phenotype relationship. OMIM is the online continuation of Dr. Victor McKusick's Mendelian Inheritance in Man (MIM) 1966-1998. MIM/OMIM is produced and curated at the Johns Hopkins University School of Medicine (JHUSOM).
| Name | OMIM | Gene | OCULOCUTANEOUS ALBINISM: Description by Wikipedia. |
| OCA1 | 203100 | TYR | OCA1 is caused by an alteration of the tyrosinase gene, and can occur in two variations. The first is OCA1a, and means that the organism cannot develop pigment at all. The hair is usually white (often translucent) and the skin very pale. Vision usually ranges from 20/200 to 20/400. The second is OCA1b, which has several subtypes itself. Some individuals with OCA1b can tan and also develop pigment in the hair. One subtype of OCA1b is called OCA1b TS (temperature sensitive), where the tyrosinase can only function below a certain temperature, which causes the body hair in cooler body regions to develop pigment (i.e. get darker). (An equivalent mutation produces the coat pattern in Siamese cats.) Another variant of OCA1b, called Albinism, yellow mutant type (OMIM: 606952) is more common among the Amish than in other populations, and results in blonde hair and the eventual development of skin pigmentation during infancy, though at birth is difficult to distinguish from other types. |
| OCA2 | 203200 | OCA2 | The most common type of albinism, is caused by mutation of the P gene. People with OCA2 generally have more pigment and better vision than those with OCA1, but cannot tan like some with OCA1b. A little pigment can develop in freckles or moles. People with OCA2 usually have fair skin but often not as pale as OCA1, and pale blonde to golden, strawberry blonde, or even brown hair, and most commonly blue eyes. Affected people of African descent usually have a different phenotype (appearance): yellow hair, pale skin, and blue, gray or hazel eyes. The gene MC1R doesn't cause OCA2, but does affect its presentation. |
| OCA3 | 203290 | TYRP1 | Has only been partially researched and documented. It is caused by mutation of the tyrosinase-related protein-1 (Tyrp1) gene. Cases have been reported in Africa and New Guinea. Affected individuals typically have red hair, reddish-brown skin and blue or gray eyes. Variants may include rufous oculocutaneous albinism (ROCA or xanthism) (OMIM: 278400). |
+
Other scientific sources |
| Oculocutaneous Albinism Type 4 (OCA4) - Because OCA2 and OCA4 are phenotypically similar, it is not possible to accurately diagnose OCA4 based on clinical findings alone. SLC45A2 (previously called MATP and AIM1) is the only gene in which mutation is known to cause OCA4. |
| Oculocutaneous Albinism Type 5 (OCA5) - A U.S. National Library of Medicine National Institutes of Health Study: OCA5, a novel locus for non-syndromic oculocutaneous albinism, maps to chromosome 4q24. by Kausar T, Bhatti MA, Ali M, Shaikh RS, Ahmed ZM. |
| Oculocutaneous Albinism Type 6 (OCA6) - Oculocutaneous albinism type 6 (OCA6) is a type of oculocutaneous albinism characterized by light hair at birth that darkens with age, white skin, transparent irides, photophobia, nystagmus, foveal hypoplasia and reduced visual acuity and that is due to mutations in the SLC24A5 gene. |
| Oculocutaneous Albinism Type 7 (OCA7) - OCA7 is due to a mutation in the C10orf11 gene (10q22.3) encoding a 198 amino acid protein. Currently, little is known about the biological function of this gene in humans and its role in OCA7 pathogenesis. |
To be clear: contrary to what the Genetic testing for money people might tell you: Y-dna and Mtdna cannot tell you anything about Race. (To wit): Blacks and Whites have EXACTLY THE SAME GENES! The only difference between a normal Black person and a White Albino person, is the "Non-Working" (i.e. Recessive) condition of the following Melanin producing genes.
Question: What is a Recessive Gene?
United States National Human Genome Research Institute
NHGRI is located on the National Institutes of Health (NIH) campus in Bethesda, MD.
Recessive (gene) is a quality found in the relationship between two versions of a gene. Individuals receive one version of a gene, called an allele, from each parent. If the alleles are different, the dominant allele will be expressed, while the effect of the other allele, called recessive, is masked. In the case of a recessive genetic disorder, (like Albinism), an individual must inherit two copies of the mutated allele in order for the disease to be present.
The Tyrosinase locus (TYR) of Oculocutaneous albinism type 1 (OCA1) has been mapped to chromosome 11, region q14 >> q21
Oculocutaneous Albinism type 2 (OCA2) (P gene) has been mapped to chromosome 15 between positions 12 and 13.1
Oculocutaneous Albinism type 3 (OCA3) (Tyrp1) has been mapped chromosome 9 at position 23
Oculocutaneous Albinism type 4 (OCA4) (SLC45A2) has been mapped to chromosome 5 at position 13.2
Oculocutaneous Albinism type 5 (OCA5) (non-syndromic Oculocutaneous Albinism) has been mapped to chromosome 4 at position 24
Oculocutaneous Albinism type 6 (OCA6) (SLC24A5) has been mapped to chromosome 15 at position 21.1
Oculocutaneous Albinism type 7 (OCA7) (C10orf11 Gene) (Encoding a 198 amino acid protein) has been mapped to chromosome 10 at position 22.3.
National Organization for Albinism and Hypopigmentation
Information Bulletin - What is Albinism?
Albinism is an inherited genetic condition that reduces the amount of melanin pigment formed in the skin, hair and/or eyes. Albinism occurs in all racial and ethnic groups throughout the world. In the U.S., approximately one in 18,000 to 20,000 people has some type of albinism. In other parts of the world, the occurrence can be as high as one in 3,000. Most children with albinism are born to parents who have normal hair and eye color for their ethnic backgrounds.
A common myth is that people with albinism have red eyes. Although lighting conditions can allow the blood vessels at the back of the eye to be seen, which can cause the eyes to look reddish or violet, most people with albinism have blue eyes, and some have hazel or brown eyes. There are different types of albinism and the amount of pigment in the eyes varies; however, vision problems are associated with albinism. [unsaid - but NOT automatic to Albinism].
The last statement is very important because how many time have you heard a European say that they are not Albinos because they DON'T have Bad eyesight/Red eyes?
Please remember that these studies and conclusions are put forth mostly by Albino derived people,
who are mostly in denial about their condition. Thus their conclusions will always skirt true reasoning.
Melanin is produced by the oxidation of the amino acid tyrosine, followed by polymerization. The pigment is produced in a specialized group of cells known as melanocytes.
In the skin, melanogenesis occurs after exposure to UV radiation, causing the skin to visibly tan. Melanin is an effective absorber of light; the pigment is able to dissipate over 99.9% of absorbed UV radiation. Because of this property, melanin is thought to protect skin cells from UVB radiation damage, reducing the risk of cancer. Furthermore, though exposure to UV radiation is associated with increased risk of malignant melanoma, a cancer of the melanocytes, studies have shown a lower incidence for skin cancer in individuals with more concentrated melanin, i.e. darker skin tone. Nonetheless, the relationship between skin pigmentation and photoprotection is still being clarified.
In humans, melanin is the primary determinant of skin color. It is also found in hair, the pigmented tissue underlying the iris of the eye, and the stria vascularis of the inner ear. In the brain, tissues with melanin include the medulla and pigment-bearing neurons within areas of the brainstem, such as the locus coeruleus and the substantia nigra. It also occurs in the zona reticularis of the adrenal gland.
The melanin in the skin is produced by melanocytes, which are found in the basal layer of the epidermis. Although, in general, human beings possess a similar concentration of melanocytes in their skin, the melanocytes in some individuals and ethnic groups produce variable amounts of melanin. Some humans have very little or no melanin synthesis in their bodies, a condition known as albinism.
Because melanin is an aggregate of smaller component molecules, there are many different types of melanin with differing proportions and bonding patterns of these component molecules. Both pheomelanin and eumelanin are found in human skin and hair, but eumelanin is the most abundant melanin in humans, as well as the form most likely to be deficient in albinism.
Melanin in the eyes, in the iris and choroid, helps protect them from ultraviolet and high-frequency visible light; people with gray, blue, and green eyes are more at risk for sun-related eye problems. Further, the ocular lens yellows with age, providing added protection. However, the lens also becomes more rigid with age, losing most of its accommodation — the ability to change shape to focus from far to near — a detriment due probably to protein crosslinking caused by UV exposure.
Recent research suggests that melanin may serve a protective role other than photoprotection. Melanin is able to effectively ligate metal ions through its carboxylate and phenolic hydroxyl groups, in many cases much more efficiently than the powerful chelating ligand ethylenediaminetetraacetate (EDTA). Thus, it may serve to sequester potentially toxic metal ions, protecting the rest of the cell. This hypothesis is supported by the fact that the loss of neuromelanin observed in Parkinson's disease is accompanied by an increase in iron levels in the brain.
Sean Dunne @irishcentral May 27,2015 04:00 AM
Have you ever wondered where the Irish get their light skin color from? Well, it appears we may now have the answer. A major new US study at Penn State University has found that Europeans' light skin stems from a gene mutation from a single person who lived 10,000 years ago. Scientists made the discovery after identifying a key gene that contributes to lighter skin color in Europeans, and the Irish fall into this category.
The Mail Online reports that, in earlier research, Keith Cheng from Penn State College of Medicine reported that one amino acid difference in the gene SLC24A5 is a key contributor to the skin color difference between Europeans and West Africans. This is undoubtedly where the Irish get their light skin from. "The mutation in SLC24A5 changes just one building block in the protein, and contributes about a third of the visually striking differences in skin tone between peoples of African and European ancestry," he said.
Cheng and his team studied segments of genetic code that have a mutation and are located closely on the same chromosome and are often inherited together. The mutation, called A111T, is found in virtually everyone of European ancestry. A111T is also found in populations in the Middle East and Indian subcontinent, but not in high numbers in Africans.
All individuals from the Middle East, North Africa, East Africa and South India who carry the A111T mutation share traces of the ancestral genetic code. According to the researchers, this indicates that all existing instances of this mutation originate from the same person. The pattern of people with this lighter skin color mutation suggests that the A111T mutation occurred somewhere between the Middle East and the Indian subcontinent.
 |
 |
 |
 |
 |
 |
TYR Gene - The official name of this gene is “tyrosinase.”
How are changes in the TYR gene related to health conditions?
oculocutaneous albinism - caused by mutations in the TYR gene
More than 100 mutations in the TYR gene have been identified in people with oculocutaneous albinism type 1. These mutations disrupt the normal production of melanin, which reduces coloring of the hair, skin, and eyes and causes problems with vision. Most TYR mutations eliminate the activity of tyrosinase, preventing melanocytes from producing any melanin throughout life. These mutations cause a form of oculocutaneous albinism called type 1A (OCA1A). People with this form of albinism have white hair, light-colored eyes, and very pale skin that does not tan. Other mutations in the TYR gene reduce but do not eliminate tyrosinase activity. These mutations, which allow some melanin to be produced, cause oculocutaneous albinism type 1B (OCA1B). People with type 1B are also born with white hair, light-colored eyes, and pale skin, but hair and eye color often darken over time and skin may tan.
Where is the TYR gene located?
Cytogenetic Location: 11q14.3
Molecular Location on chromosome 11: base pairs 89,177,871 to 89,295,758
.
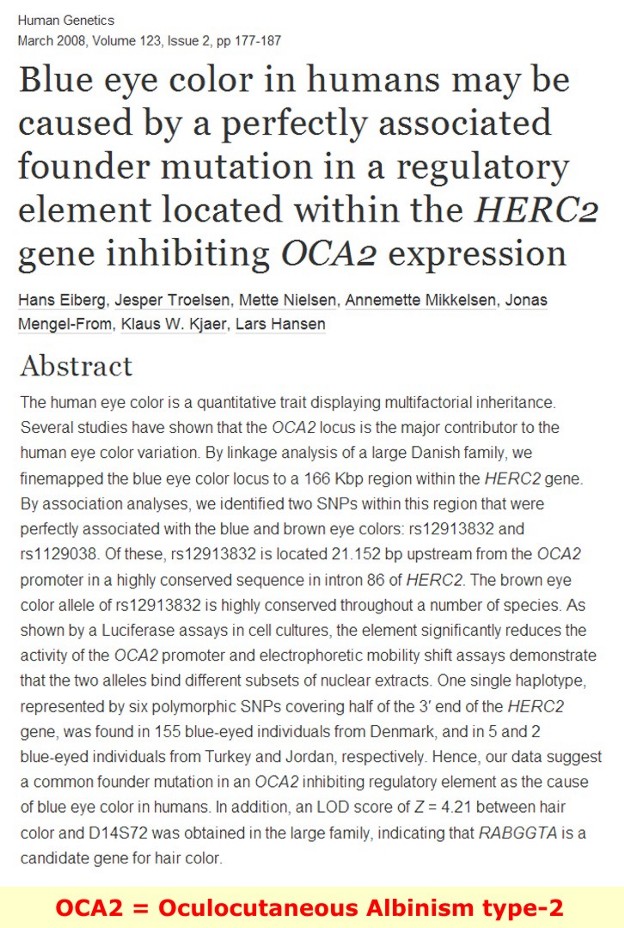
OCA2 gene (formerly called the P gene) - The official name of this gene is “oculocutaneous albinism II.”
How are changes in the OCA2 gene related to health conditions?
Oculocutaneous albinism - caused by mutations in the OCA2 gene
More than 80 mutations in the OCA2 gene have been identified in people with oculocutaneous albinism type 2. People with this form of albinism often have light yellow, blond, or light brown hair; creamy white skin; light-colored eyes; and problems with vision. The most common OCA2 mutation is a large deletion in the gene, which is found in many affected individuals of sub-Saharan African heritage. Other OCA2 gene mutations, including changes in single DNA building blocks (base pairs) and small deletions, are more common in other populations. Mutations in the OCA2 gene disrupt the normal production of melanin, which reduces coloring of the hair, skin, and eyes and affects vision.
Where is the OCA2 gene located?
Cytogenetic Location: 15q
Molecular Location on chromosome 15: base pairs 27,754,874 to 28,099,336
 |
 |
TYRP1 gene - The official name of this gene is “tyrosinase-related protein 1.”
How are changes in the TYRP1 gene related to health conditions?
oculocutaneous albinism - caused by mutations in the TYRP1 gene
A small number of mutations in the TYRP1 gene have been found to cause oculocutaneous albinism type 3. This condition includes a form of albinism called rufous oculocutaneous albinism, which has been described primarily in dark-skinned people from southern Africa. Affected individuals have reddish-brown skin, ginger or red hair, and hazel or brown irises. Two TYRP1 mutations are known to cause this form of albinism in individuals from Africa. One mutation replaces a protein building block (amino acid) in tyrosine-related protein 1 with a signal that prematurely stops protein production. This mutation, written as Ser166Ter or S166X, affects the amino acid serine at protein position 166. The other mutation, written as 368delA, deletes a single DNA building block from the TYRP1 gene. Other alterations in this gene have been reported in a few affected people of non-African heritage. Most TYRP1 mutations lead to the production of an abnormally short, nonfunctional version of tyrosinase-related protein 1. Because this enzyme plays a role in normal pigmentation, its loss leads to the changes in skin, hair, and eye coloration that are characteristic of oculocutaneous albinism.
Where is the TYRP1 gene located?
Cytogenetic Location: 9p23
Molecular Location on chromosome 9: base pairs 12,693,384 to 12,710,284
.
SLC45A2 gene - The official name of this gene is “solute carrier family 45, member 2.”
How are changes in the SLC45A2 gene related to health conditions?
oculocutaneous albinism - caused by mutations in the SLC45A2 gene
More than 20 mutations in the SLC45A2 gene are responsible for oculocutaneous albinism type 4. The most common SLC45A2 mutation in the Japanese population switches a single protein building block (amino acid) in the SLC45A2 protein. Specifically, this mutation replaces the amino acid aspartic acid with the amino acid asparagine at protein position 157 (written as Asp157Asn or D157N). Other mutations, including changes in single amino acids and deletions or insertions of genetic material in the SLC45A2 gene, have also been reported in several populations worldwide. Mutations in this gene reduce or eliminate the function of the SLC45A2 protein in melanin production. Because this protein is important for normal pigmentation, its loss leads to changes in skin, hair, and eye coloration and problems with vision that are characteristic of oculocutaneous albinism type 4.
Where is the SLC45A2 gene located?
Cytogenetic Location: 5p13.2
Molecular Location on chromosome 5: base pairs 33,944,615 to 33,984,674
.
Please keep in mind that there are at LEAST 3 more types of Oculocutaneous Albinism: type OCA5, OCA6, OCA7, and more are expected to be discovered.
The official name of this gene is “melanocortin 1 receptor (alpha melanocyte stimulating hormone receptor).”
What is the normal function of the MC1R gene?
The MC1R gene provides instructions for making a protein called the melanocortin 1 receptor. This receptor plays an important role in normal pigmentation. The receptor is primarily located on the surface of melanocytes, which are specialized cells that produce a pigment called melanin. Melanin is the substance that gives skin, hair, and eyes their color. Melanin is also found in the light-sensitive tissue at the back of the eye (the retina), where it plays a role in normal vision.
Melanocytes make two forms of melanin, eumelanin and pheomelanin. The relative amounts of these two pigments help determine the color of a person's hair and skin. People who produce mostly eumelanin tend to have brown or black hair and dark skin that tans easily. Eumelanin also protects skin from damage caused by ultraviolet (UV) radiation in sunlight. People who produce mostly pheomelanin tend to have red or blond hair, freckles, and light-colored skin that tans poorly. Because pheomelanin does not protect skin from UV radiation, people with more pheomelanin have an increased risk of skin damage caused by sun exposure.
Common variations (polymorphisms) in the MC1R gene are associated with normal differences in skin and hair color. Certain genetic variations are most common in people with red hair, fair skin, freckles, and an increased sensitivity to sun exposure. These MC1R polymorphisms reduce the ability of the melanocortin 1 receptor to stimulate eumelanin production, causing melanocytes to make mostly pheomelanin. Although MC1R is a key gene in normal human pigmentation, researchers believe that the effects of other genes also contribute to a person's hair and skin coloring. The melanocortin 1 receptor is also active in cells other than melanocytes, including cells involved in the body's immune and inflammatory responses. The receptor's function in these cells is unknown.
Does the MC1R gene share characteristics with other genes?
The MC1R gene belongs to a family of genes called GPCR (G protein-coupled receptors).
A gene family is a group of genes that share important characteristics. Classifying individual genes into families helps researchers describe how genes are related to each other. For more information, see What are gene families? in the Handbook.
Cancers - increased risk from variations of the MC1R gene
Many genetic changes in the MC1R gene increase the risk of developing skin cancer, including a common, serious form of skin cancer that begins in melanocytes (melanoma). Alterations in the MC1R gene disrupt the ability of the melanocortin 1 receptor to trigger eumelanin production in melanocytes. Because eumelanin normally protects skin from the harmful effects of UV radiation, a lack of this pigment leaves fair skin more vulnerable to damage from sun exposure. Skin damage caused by UV radiation from the sun is a major risk factor for developing melanoma and other forms of skin cancer.
Studies suggest that variations in the MC1R gene may also increase the risk of developing melanoma in the absence of UV radiation-related skin damage. In these cases, melanomas can occur in people of dark or light skin coloring. These cancers are often associated with mutations in additional genes related to melanoma risk, such as the BRAF and CDKN2A genes. Researchers are working to explain the complex relationship among MC1R variations, other genetic and environmental factors, and melanoma risk.
Oculocutaneous albinism - course of condition modified by mutations in the MC1R gene
Certain genetic changes in the MC1R gene modify the appearance of people with oculocutaneous albinism type 2. This form of albinism, which is caused by mutations in the OCA2 gene, is characterized by fair hair, light-colored eyes, creamy white skin, and vision problems. People with genetic changes in both the OCA2 and MC1R genes have many of the usual features of oculocutaneous albinism type 2; however, they typically have red hair instead of the usual yellow, blond, or light brown hair seen with this condition.
 |
 |
Wiki:
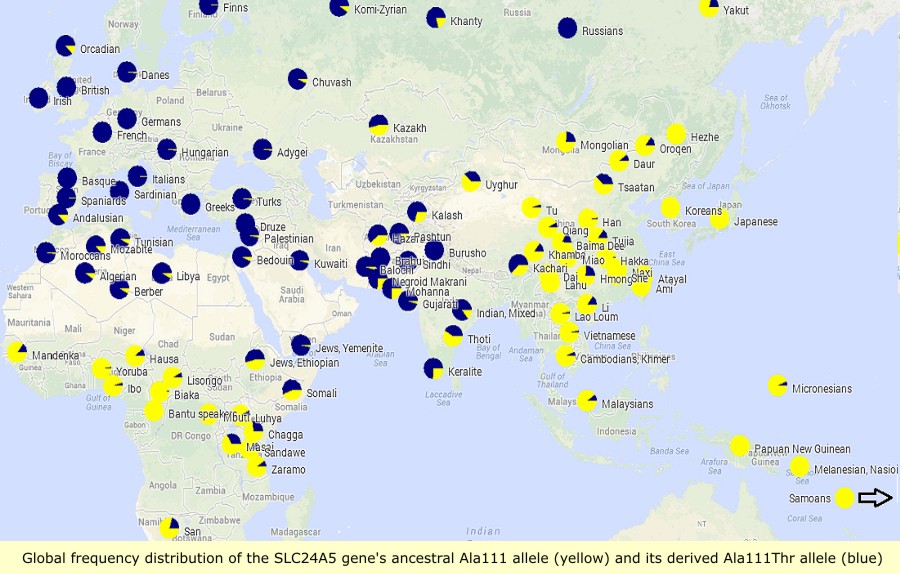
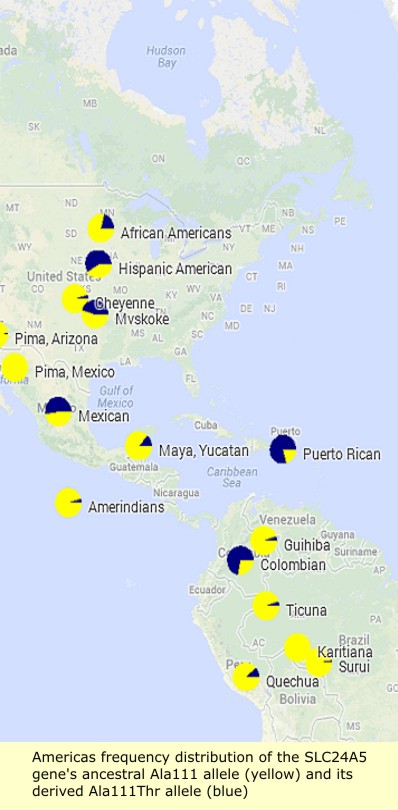
Sodium/potassium/calcium exchanger 5 (NCKX5), also known as solute carrier family 24 member 5 (SLC24A5), is a protein that in humans is encoded by the SLC24A5 gene that has a major influence on natural skin colour variation. The NCKX5 protein is a member of the potassium-dependent sodium/calcium exchanger family. Sequence variation in the SLC24A5 gene, particularly a non-synonymous SNP changing the amino acid at position 111 in NCKX5 from alanine to threonine, has been associated with differences in skin pigmentation.
The SLC24A5 gene's derived threonine or Ala111Thr allele (rs1426654[3]) has been shown to be a major factor in the light skin tone of Europeans compared to Africans, and is believed to represent as much as 25–40% of the average skin tone difference between Europeans and West Africans. It has been the subject of recent selection in Western Eurasia, and is fixed in European populations.
The SLC24A5 gene, in humans, is located on the long (q) arm of chromosome 15 on position 21.1, from base pair 46,200,461 to base pair 46,221,881.
NCKX5 is 43 kDa protein that is partially localized to the trans-Golgi network in melanocytes. Removal of the NCKX5 protein disrupts melanogenesis in human and mouse melanocytes, causing a significant reduction in melanin pigment production. Site-directed mutagenesis corresponding to a non-synonymous single nucleotide polymorphism in SLC24A5 alters a residue in NCKX5 (A111T) that is important for NCKX5 sodium-calcium exchanger activity.
SLC24A5 appears to have played a key role in the evolution of light skin in humans of European ancestry. The gene's function in pigmentation was discovered in zebrafish as a result of the positional cloning of the gene responsible for the "golden" variety of this common pet store fish. Evidence in the International HapMap Project database of genetic variation in human populations showed that Europeans, represented by the "CEU" population, had two primary alleles differing by only one nucleotide, changing the 111th amino acid from alanine to threonine, abbreviated "A111T"
The derived threonine allele (Ala111Thr; also known as A111T or Thr111) represented 98.7 to 100% of the alleles in European samples, while the ancestral or alanine form was found in 93 to 100% of samples of Sub-Saharan Africans, East Asians and Indigenous Americans. The variation is a SNP polymorphism rs1426654, which had been previously shown to be second among 3011 tabulated SNPs ranked as ancestry-informative markers. This single change in SLC24A5 explains between 25 and 38% of the difference in skin melanin index between peoples of West African vs. European Ancestry.
Furthermore, the European mutation is associated with the largest region of diminished genetic variation in the CEU HapMap population, suggesting the possibility that the A111T mutation may be the subject of the single largest degree of selection in human populations of European ancestry. It is theorised that selection for the derived allele is based on the need for sunlight to produce the essential nutrient vitamin D. In northerly latitudes, where there is less sun, greater requirement for body coverage due to colder climate, and frequently, diets poor in vitamin D, making lighter skin more suitable for survival. Tests for this variation have obvious application to forensic science.
It has been estimated that the threonine allele became predominant among Europeans 11,000 to 19,000 years ago.
 |
 |
 |
 |
 |
 |
 |
 |
Always Remember THIS: |
Often we read racial population analyses that attempt to differentiate between Blacks and Whites genetically by haplogroups. And we always ask: "How Can They Do That, When There is No such A Thing As White Genes:" (whatever genes Whites have, Black have, and had them first, because of course Whites are the Albinos of Blacks).
Thus the reason we feature the following study is not because of the accuracy or import of it's subject matter, rather it is because of it's method. It is one of the few studies which admits by inference, that a humans level of Whiteness equates with his level of Albinism. Thus, the ONLY TRUE MEASURE OF WHITE/ALBINISM IS MELANIN CONTENT: i.e Melanin Index. Thus, "DARK" Europeans as those Europeans with significant Black admixture - AFTER THE ONSET OF THE ANCIENT ALBINISM EVENT.
Abstract
We carried out an admixture analysis of a sample comprising 1,019 individuals from all the provinces of Cuba. We used a panel of 128 autosomal Ancestry Informative Markers (AIMs) to estimate the admixture proportions. We also characterized a number of haplogroup diagnostic markers in the mtDNA and Y-chromosome in order to evaluate admixture using uniparental markers. Finally, we analyzed the association of 16 single nucleotide polymorphisms (SNPs) with quantitative estimates of skin pigmentation. In the total sample, the average European, African and Native American contributions as estimated from autosomal AIMs were 72%, 20% and 8%, respectively. The Eastern provinces of Cuba showed relatively higher African and Native American contributions than the Western provinces. In particular, the highest proportion of African ancestry was observed in the provinces of Guantanamo (40%) and Santiago de Cuba (39%), and the highest proportion of Native American ancestry in Granma (15%), Holgum (12%) and Las Tunas (12%). We found evidence of substantial population stratification in the current Cuban population, emphasizing the need to control for the effects of population stratification in association studies including individuals from Cuba. The results of the analyses of uniparental markers were concordant with those observed in the autosomes. These geographic patterns in admixture proportions are fully consistent with historical and archaeological information. Additionally, we identified a sex-biased pattern in the process of gene flow, with a substantially higher European contribution from the paternal side, and higher Native American and African contributions from the maternal side. This sex-biased contribution was particularly evident for Native American ancestry. Finally, we observed that SNPs located in the genes SLC24A5 and SLC45A2 are strongly associated with melanin levels in the sample.
The program ADMIXMAP was used to run a linear regression analysis conditioning on individual ancestry. The P-values for 86 AIMs unlinked to the pigmentation markers were used to estimate the lambda inflation factor. We report the P-values after Genome Control (GC) correction. doi:l 0.1 371 /journal.pgen.1 004488.1002 SLC24A3 gene {P= 1.2x10"-'), rsl6891982 and rs35395 located on the SLC45A2 (MATP) gene (P=1.7xl0"2" and P = 2.8xl0"", respectively), and rsl2913832 located on the HERC2 gene, linked to 0CA2 (P = 0.0018). In order to evaluate if there was evidence of residual stratification unaccounted for in the analysis based on the three-parental model, we used the P-values obtained for 86 AIMs located more than 5 cM apart from the 16 pigmentation markers to estimate the lambda inflation factor. We observed evidence of residual stratification (lambda = 1.38). Therefore, we implemented genome control (GC) [38] methods to correct for type I error inflation. After GC-correction, SLC24A5 rsl426654 (P = 5.1 x 10"'''), SLC45A2 rsl6891982 (P = 2.9x10"") and SLC45A2 rs35395 (P= 1.5x10"") remained significant after Bonferroni correction. However, the P-value for HERC2 rsl2913832 (P = 0.0078) slightly exceeded the Bonferroni-corrected threshold. Table 2 reports the GC-corrected P- values for all the pigmentation markers. Assuming an additive model, we estimated that each copy of rsl 426654 allele A and rsl6891982 allele G decrease the melanin index by 5.04 and 3.40 units, respectively. The HERC2 SNP rsl 29 13832 has a substantially smaller effect, with each copy of the G allele, which has been associated with blue iris color in previous studies [25-27], decreasing melanin index by approximately 1.11 units. Finally, we repeated the analysis including the genotypes of rsl 426654, rsl 689 1982 and rsl 29 13832 as covariates. This analysis showed that the P-value observed for rs35395 at the SLC45A2 locus was no longer significant, indicating that the significant result for this marker is primarily due to its linkage with rsl 689 1982, which is located approximately 3 kb apart from rs35395 on chromosome 5. None of the other 12 SNPs surveyed had significant effects on melanin levels after conditioning for the rsl 426654, rsl 689 1982 and rsl 29 13832 polymorphisms.
Discussion
Here we report an analysis of the admixture proportions in a large sample from Cuba using a combination of highly informative AIMs, mtSNPs and Y-SNPs. One of the major strengths of this study is the careful selection of the sample, which represents aU the provinces of Cuba. The sample comprises individuals from more than 8 1 % of the Cuban municipalities, and the proportions according to province, age group, and rural/ urban population are very similar to the proportions reported in the Cuban 2002 census [39]. The distributions of gender and census categories ("bianco", "mestizo" and "negro") are slightly different from the reported 2002 census proportions. The proportion of females in the sample (58%) is higher than that reported in the census (50%). This is related to the fact that when the households were visited, relatively more women were the only household members present during the visit. With respect to the census categories, the sample included relatively more individuals classified as "mestizos" and less individuals classified as "blancos" than in the 2002 census (mestizos: 33% vs. 25%, blancos: 55% vs 65%), and the proportion of individuals classified as "negro" was overly similar in the sample and 2002 census (12% vs. 10%). There were also sKght differences in the way that the census categories were obtained: In the 2002 census, the "color" categories were classified by the census collectors, and when an individual was not present in the household, census categories were reported by family members. In the present sample, the census categories were obtained in two ways: self-reported and reported by a trained researcher and we observed a very high concordance between the two classifications.
The use of autosomal and maternally and paternally inherited polymorphisms allowed us to carry out a detailed analysis of admixture in Cuba, and the analysis by provinces identified very clear and consistent patterns. Using autosomal AIMs, we observed that the average European, African and Native American proportions in the sample were 72% (SD: ±22,61), 20% (SD: ±22,66) and 8% (SD: ±6,86), respectively. However, the amount of European ancestry tends to be higher in the Western provinces of Cuba than in the Eastern provinces. In contrast, the highest African proportions are observed in the eastern provinces of Santiago de Cuba and Guantanamo and the highest Native American contributions in the Eastern provinces of Las Tunas, Granma and Holguin. Importantly, the results based on analyses of the mtDNA and Y-chromosome SNPs are fuUy consistent with this picture. The highest Eurasian proportions observed for both the mtDNA and the Y chromosome are found in the Western provinces, particularly Matanzas and Pinar del Rio, and the highest African contributions are present in Santiago de Cuba. Regarding the Native American contribution, the mtDNA analysis also indicates that the highest Native American proportions are present in the provinces of Holguin and Las Tunas. We only observed two Native American Q;M3 haplogroups in the male sample, corresponding to individuals from the provinces of Camagiiey (in the Central region of the island) and Santiago de Cuba (in the east).
Our analyses indicate that the geographic trends observed in ancestry proportions are due, at least to some extent, to differences in the relative proportions of individuals reporting to be "bianco", "mestizo" or "negro" across provinces. The provinces of Guantanamo and Santiago de Cuba, which show the highest average African ancestry and melanin index levels, also have the highest proportion of individuals self-reporting to be "mestizo" and "negro". However, this does not seem to be the only reason behind these differences. We also observe that there are some differences in average admixture proportions within each census group between provinces. For example, the average African ancestry of individuals self-reporting to be "bianco" tends to be higher in Guantanamo, Santiago de Cuba and Granma than in other provinces. In general, our study highlights the subjectivity involved in the categories "bianco", "mestizo" and "negro". Although there are significant differences in melanin index between the three categories, there is some overlap in melanin values between these groups. This means that two individuals with the same melanin index values may report different census categ()ri(\s (e.g. "bianco" or "mestizo"). In addition to the analyses b)' j)r()\in(x\ wc also evaluated the distribution of ancestry proportions in urban vs. rural areas. We observed that the African ancestry proportions were significantly higher in urban than rural areas, and conversely, the Native American ancestry proportions were significantly higher in rural than urban areas. The geographic patterns observed in the distribution of admixture proportions are in agreement with historical and archaeological data. It is known that at the arrival of the Spaniards to Cuba, the Taino primarily inhabited the eastern regions of Cuba.
Estimates of the population distribution in the year 1510 indicate that more than 50% of the indigenous Cuban population lived in the eastern region (from Las Tunas to Guantanamo), less than 40% lived in Camagiiey and Las Villas (both in the central region of Cuba), and less than 10% inhabited the western region of the island. Within the eastern region, Holguin was the most populated area, followed by the region of Bayamo (currendy the province of Granma) [40] . The results of our study, which reveal that the province of Ilolguin has some of the highest autosomal and mtDNA Native American proportions in Cuba, are therefore in agreement with the historical sources described above and the high concentration of Taino archaeological sites in this area [41]. Historical reports indicate that the indigenous population collapsed from more than 100,000 at the arrival of the Europeans to 2,000-3,000 in 1556, primarily due to the harsh conditions of forced labor, the disruption of the agricultural system and the epidemic diseases brought by the Europeans [39,42].
In the early stages of colonization there was immigration from Europe, primarily from the Iberian Peninsula and the Canary Islands, and enslaved Africans were also brought to the island. Initially, the number of enslaved Africans was small, but increased substantially in the final period of the 1 8''^ century [40] . Although the eastern region was, at the arrival of the Europeans, the most populated region of the island, this was the region that took the longest to repopulate after the demographic collapse that occurred during the first stages of colonization. The western region had an important number of enslaved Africans working in the sugar plantations, but this was also the rcgicm that received most of the immigrants from the Iberian Peninsula [40]. In contrast to the western region, where most of the enslaved Africans came directly from Africa, in the eastern region many of the individuals of African ancestry came from Jamaica and Haiti, and were forced to work in coffee and sugar plantations. Historical sources indicate that in 1830 there were more than 50,000 enslaved Africans in Santiago de Cuba and Guantanamo, the regions where we have identified the highest African contributions [43,44]. The comparison of the relative autosomal, paternal and maternal admixture proportions clearly show that the process of admixture in Cuba has been sex-biased, with a relatively higher European contribution observed for the paternal lineages, and a higher African and Native American contribution in the maternal lineages. This sex-biased contribution is particularly evident for the Native American ancestry. We estimated the average maternal Native American proportion to be 34.5% in the sample, in sharp contrast to the autosomal (8%) and paternal (0.5%) proportions.
The African maternal, autosomal and paternal proportions were estimated to be 39%, 20% and 18%, respectively. Overall, our results are very similar to those obtained in an independent study that analyzed mtDNA and Y-chromosome variation in a Cuban sample comprising 245 individuals [9]. In this study, the authors reported that 4:5% of the mtDNA lineages were of African ancestry and 33% of Native American ancestry. In contrast, only 20% of the Y-chromosome lineages were of African ancestry, and the authors did not find any Y-chromosome Native American lineages. Thus, the genetic data confirms historical information indicating that most of the European migrants to Cuba were males, and that the process of mixing primarily took place between European males and Native American females, during the first stages of colonization, and African females during the slave trade period [5,9,12,13]. We explored the relationship between admixture estimates based on genetic markers, melanin levels measured with a reflectometer, and self-reported census categories ("bianco", "mestizo" and "negro"). We observed strong relationships between admixture proportions and melanin levels, admixture proportions and census categories, and melanin levels and census categories (see Results section).
Overall, these analyses show that there is very substantial population stratification in the current Cuban population, both across and edso within self-reported census categories, emphasizing the need to control for the effects of population stratification in association studies in this population. A clear example of the consequences of stratification can be seen in an analysis of the results of a linear regression model without conditioning for individual ancestry proportions, based on 86 ATMs that are located more than 5 cM apart from any of the pigmentation markers analyzed in this study. In such analysis, 64 of the 86 AIMs (74.4%) surpass the Bonferroni-corrected significance threshold (P = 5.8x10 *). In contrast, none of the AIMs surpass this threshold when the analysis is carried out conditioning on individual ancestry. This implies that in case- control studies in which there are differences in ancestry proportions between the case and control group, or association analysis of quantitative traits that have different distributions in the parental populations, such as pigmentation, there would be a dramatic inflation in the number of false positives.
We observed that even after conditioning for individual ancestry - there was evidence of residual stratification in the Cuban sample, although of relatively small magnitude (lambda 1.38). Consequently, the P- values observed for the pigmentation markers were corrected using Genome Control (GC) strategies. Finally, we also evaluated the association of 16 SNPs located within or nearby pigmentation genes with melanin levels (e.g. melanin index). These polymorphisms have been associated with pigmentary phenotypes in previous studies [21-37]. Our analysis confirms previously reported associations of rs 1426654, located within tie SLC24A5 gene (P = 5.1 xlO"'^) and rsl6891982, located within tie SLC45A2 [MATP] gene (P = 2.9x10"'^) with skin pigmentation. These two markers have the strongest effects on melanin levels described in human populations, and in our study we estimated that each copy of rs 1426654 allele A and rs 1689 1982 allele G decrease the melanin index by 5.04 and 3.40 units, respectively. The marker rsl2913832, which is located within the HERC2 gene and is known to affect the transcription of the 0CA2 gene, showed a significant effect in the initial ADMIXMAP association tests (P = 0.0018), but it did not surpass the Bonferroni corrected threshold (P 0.0033) after CG-correction (P = 0.0077). This marker is strongly associated with blue eye color in European populations [25-27], but it has also been associated with skin pigmentation, tanning response and hair color in previous studies.
The Central Asian Albinos - todays White Europeans and unmixed Turks - are clearly loath to admit that they are Albinos. So to hide this truth, they utilize all manner of "Double-Speak": that is defining Albinism, but turning aside all inference to themselves.
They say things like: OCA2 is rare in Europe, but more common in Africa. That's such a silly lie: ALL White Europeans are ALREADY OCA2, so to hide that; they only count as Albino, those of their number who have genetic vision problems because of their OCA2 Albinism: (Another lie they tell is that ALL Albinos have vision problems).
So for a better understanding, lets DEFINE OCA2.
OCA2 stands for Oculocutaneous Albinism type II. Oculocutaneous Albinism type II is a DISEASE!!!!!
These are the symptoms of OCA2.
Genetics Home Reference
The OCA2 gene (formerly called the P gene) provides instructions for making a protein called the P protein. This protein is located in melanocytes, which are specialized cells that produce a pigment called melanin. Melanin is the substance that gives skin, hair, and eyes their color. Melanin is also found in the light-sensitive tissue at the back of the eye (the retina), where it plays a role in normal vision.
NOAH (National Organization for Albinism and Hypopigmentation).
A common myth is that people with albinism have red eyes. Although lighting conditions can allow the blood vessels at the back of the eye to be seen, which can cause the eyes to look reddish or violet, most people with albinism have blue eyes, and some have hazel or brown eyes. There are different types of albinism and the amount of pigment in the eyes varies; however, vision problems are associated with albinism.
Did you notice in the "Genetics Home Reference" definition where it said that "The OCA2 gene was formerly called the P gene"? Now why would White people RENAME a gene after a disease?
THE HUMAN BODY DOES NOT "NORMALLY" COME WITH DISEASE! So why did the Albinos RENAME the "P" gene, and give it the name of a DISEASE? They did that when they found out that the MUTATED form of the "P" gene was "NORMAL" in THEM! OCA2 "IS" the "MUTATED" FORM of the "P" gene.
To put it plainly: In a world without Albino lies: A Normal Black person’s gene would be called a "P" gene. And only the MUTATED form found in Europeans and African Albinos, would be called the OCA2 gene. Since ALL Europeans have the OCA2 gene, therefore they are all Albinos. And of course it's rare in Africa, most Africans are NOT Albinos.
In combating the Albino lies of today, it is often useful to read how they looked thousands of years ago.
In Herodotus's "History of the Persian Wars" of the dozens of peoples that he describes in the book; he chooses to describe only three peoples by racial type. The Colchians above; whom he describes as "black-skinned and have woolly hair". And the Budini of Gelonus (east-central Ukraine), whom he describes as (they have all deep blue eyes, and bright red hair)
The Roman historian Cornelius Tacitus (56-118 A.D.) said this about them: For my own part, I agree with those who think that the tribes of Germany are free from all taint of intermarriages with foreign nations, and that they appear as a distinct, unmixed race, like none but themselves. Hence, too, the same physical peculiarities throughout so vast a population. All have fierce blue eyes, red hair, huge frames, fit only for a sudden exertion. They are less able to bear laborious work. Heat and thirst they cannot in the least endure; to cold and hunger their climate and their soil inure them.
From those passages we know for sure what White Europeans looked like when they first invaded Europe - they were Pure Albinos.
But today, they rarely have the RED HAIR and BLUE EYES of their ancestors - What Happened?
THIS HAPPENED!
"Since the turn of the century, people born with blue eyes in the United States have dramatically decreased, with only about 10 percent having blue eyes today.
According to Mark Grant, an epidemiologist from Loyola University in Chicago. During the turn of the last century, the percentage of people with blue eyes stood at 57.4% for those born between 1899 through 1905; and 33.8% for those born between 1936 through 1951. According to Grant, in a study titled "Cohort effects in a genetically determined trait: eye color among US whites." This decrease in the occurrence of blue eyes is due to many factors, with the majority pointing to the increase in brown-eyed immigrants, mainly Hispanics and Asians, as well as heightened interracial relationships: as the other determinant, (when a normal Black person and a European make a baby, the baby GAINS varied ability to make MELANIN). Blue eyes, next to green, are the rarest eye color in the world, as people of counties in Asia and Africa possess brown eyes."
| Click for Realhistoryww Home Page |